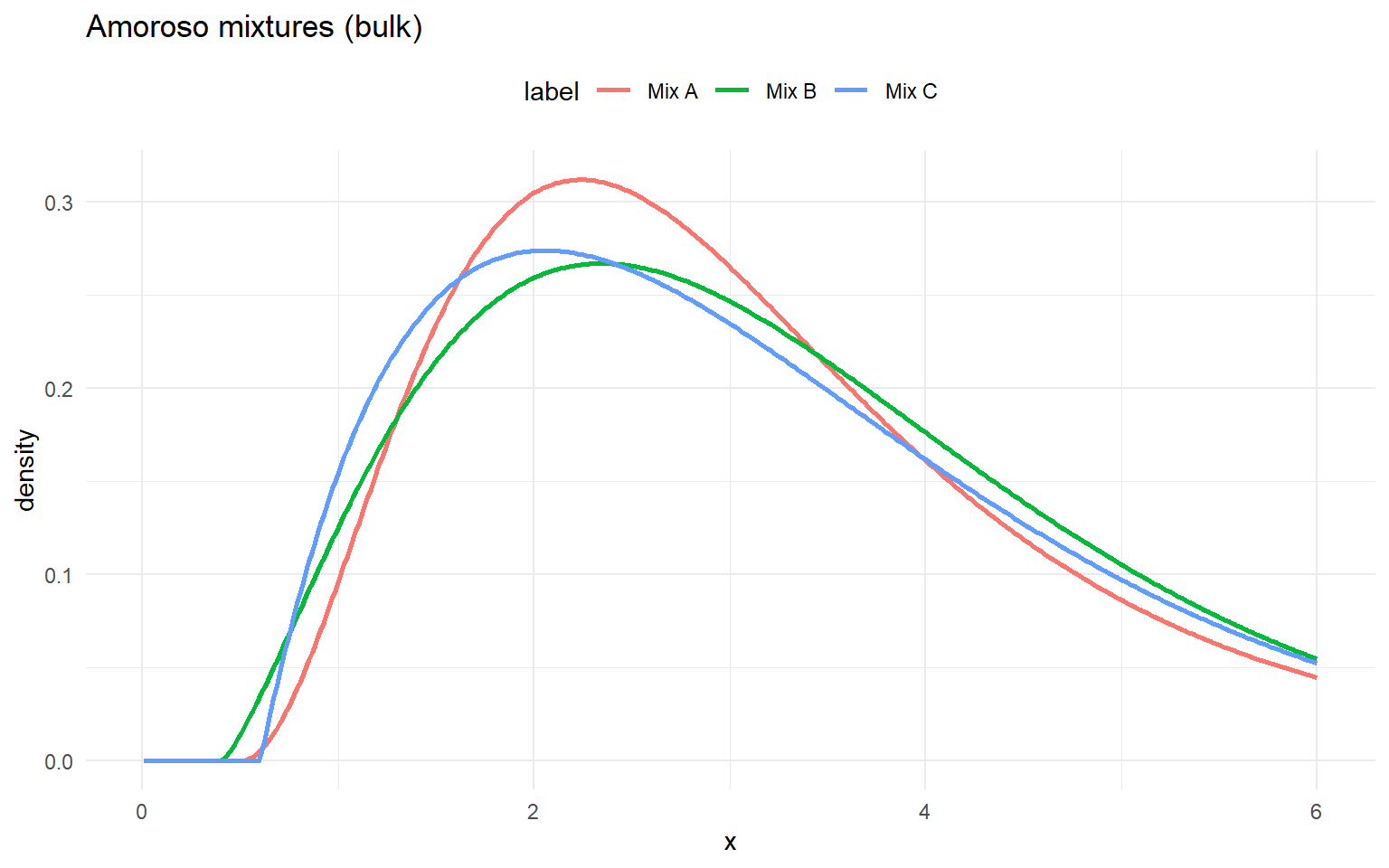
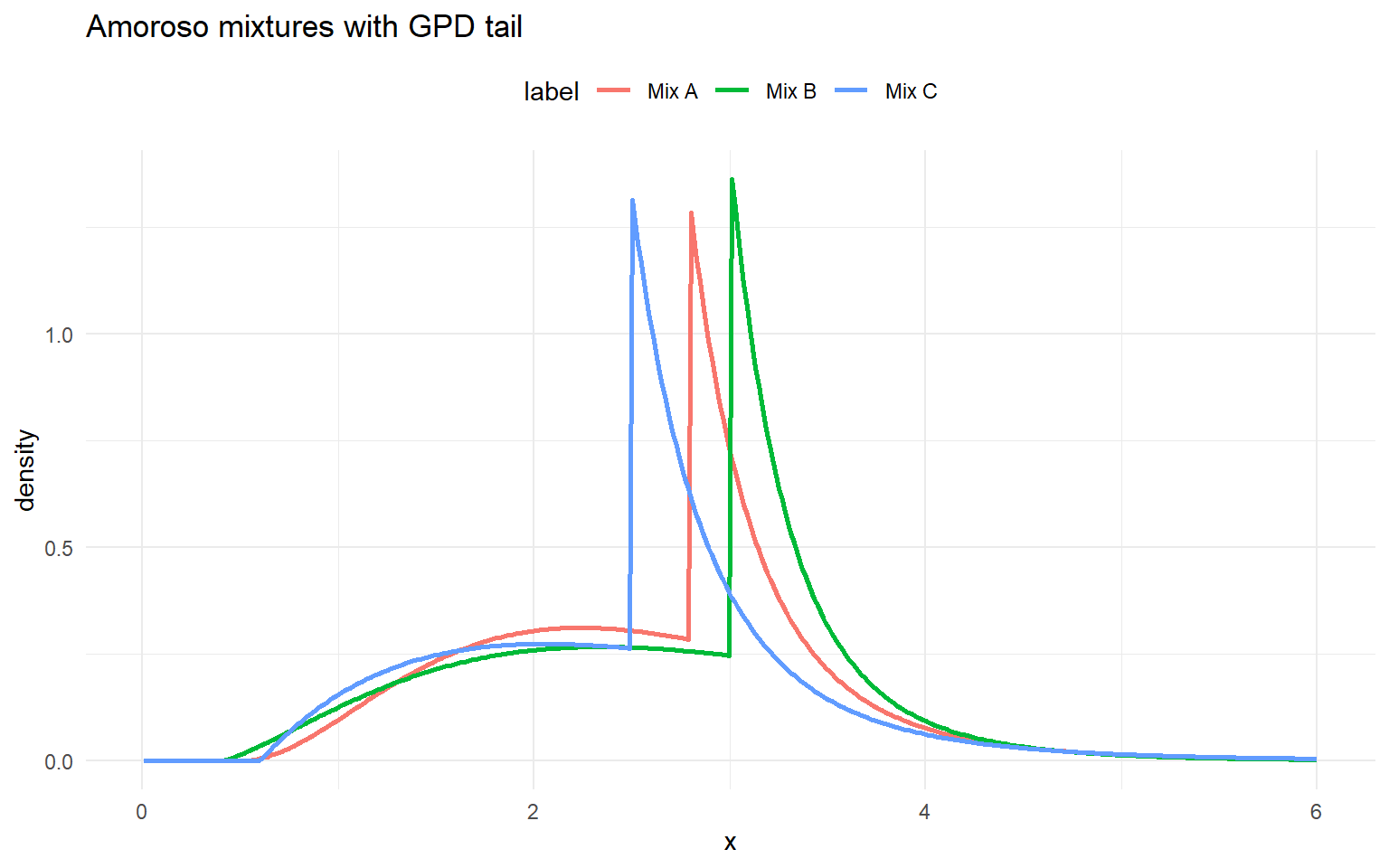
amor_gpd_sets <- list(
list(label = "Mix A", w = c(0.6, 0.3, 0.1), loc = c(0.5, 0.5, 0.5), scale = c(1.0, 1.3, 1.6), shape1 = c(2.5, 3.0, 4.0), shape2 = c(1.2, 1.2, 1.2), threshold = 2.8, tail_scale = 0.4, tail_shape = 0.2),
list(label = "Mix B", w = c(0.5, 0.3, 0.2), loc = c(0.4, 0.6, 0.6), scale = c(1.1, 1.2, 1.5), shape1 = c(2.2, 2.8, 3.8), shape2 = c(1.1, 1.2, 1.3), threshold = 3.0, tail_scale = 0.35, tail_shape = 0.18),
list(label = "Mix C", w = c(0.4, 0.35, 0.25), loc = c(0.6, 0.6, 0.6), scale = c(0.9, 1.1, 1.4), shape1 = c(2.0, 2.6, 3.5), shape2 = c(1.0, 1.1, 1.2), threshold = 2.5, tail_scale = 0.45, tail_shape = 0.22)
)
example <- amor_gpd_sets[[1]]