Website workflow note. This page reflects the current exported API and recommended wrapper-first usage. Last updated: 2026-02-19 .
For the full package narrative, see the main package vignettes (basic, unconditional, conditional, and causal).
Unconditional DPmix: Chinese Restaurant Process (CRP)
What you’ll learn
Fit a bulk-only one-arm model with dpmix() using the CRP backend.
Use the package’s S3 surface (summary(), plot(), predict()) to diagnose and extract posterior predictive summaries.
When to use this template
You have a single outcome (y) and want a flexible density model without covariates (X).
You do not need tail splicing (GPD = FALSE).
You want adaptive clustering behavior via CRP (within a finite components cap).
Goal (in one sentence)
Estimate the density of a univariate outcome (y) using a Dirichlet-process mixture with a Chinese Restaurant Process backend.
Model : \(y_i | G \sim \int K(y_i; \theta) dG(\theta)\) where \(G \sim \text{DP}(\alpha, G_0)\)
Backend : CRP with truncation at max components
Data Setup
Code
# Load pre-generated dataset: 200 observations from mixture of 3 gamma components data ("nc_pos200_k3" )<- nc_pos200_k3$ ypaste ("Sample size:" , length (y_mixed))
Code
paste ("Mean:" , mean (y_mixed))
[1] "Mean: 4.21476750434594"
Code
paste ("SD:" , sd (y_mixed))
[1] "SD: 4.10835046697183"
Code
paste ("Range:" , paste (range (y_mixed), collapse = " to " ))
[1] "Range: 0.0403111680208858 to 19.6013451514889"
Code
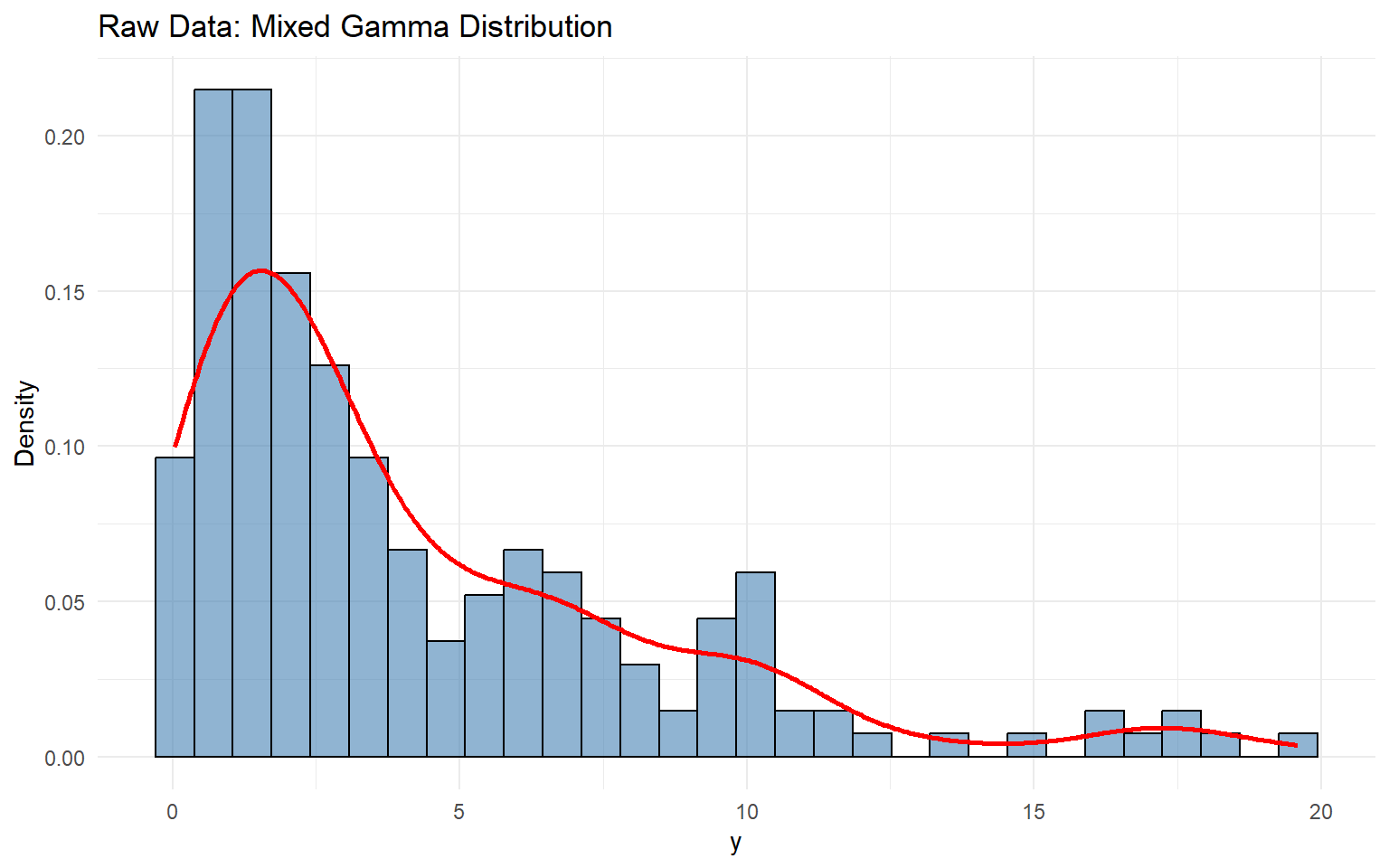
# Visualization <- data.frame (y = y_mixed)<- ggplot (df_data, aes (x = y)) + geom_histogram (aes (y = after_stat (density)), bins = 30 , alpha = 0.6 , fill = "steelblue" ,color = "black" ) + geom_density (color = "red" , linewidth = 1 ) + labs (title = "Raw Data: Mixed Gamma Distribution" , x = "y" , y = "Density" ) + theme_minimal ()print (p_raw)
Model Specification & Bundle
This page keeps the direct bundle() workflow because it is still useful for showing how the unconditional CRP specification is assembled before fitting.
Code
<- bundle (y = y_mixed,kernel = "laplace" ,backend = "crp" ,GPD = FALSE ,components = 3 ,alpha_random = TRUE ,mcmc = mcmc
Summary of MCMC Bundle
Code
CausalMixGPD bundle summary
Field Value
Backend Chinese Restaurant Process
Kernel Laplace Distribution
Components 3
N 200
X NO
GPD FALSE
Epsilon 0.025
Parameter specification
block parameter mode level prior link
meta backend info model crp
meta kernel info model laplace
meta components info model 3
meta N info model 200
meta P info model 0
concentration alpha dist scalar gamma(shape=1, rate=1)
bulk location dist component (1:3) normal(mean=0, sd=5)
bulk scale dist component (1:3) gamma(shape=2, rate=1)
notes
stochastic concentration
iid across components
iid across components
Monitors
n = 4
alpha, z[1:200], location[1:3], scale[1:3]
Running MCMC
Code
<- load_or_fit ("ex01-unconditional-dpm-crp-fit_crp" , dpmix (bundle_crp))
Summary of Fitted MCMC model
Code
MixGPD summary | backend: Chinese Restaurant Process | kernel: Laplace Distribution | GPD tail: FALSE | epsilon: 0.025
n = 200 | components = 3
Summary
Initial components: 3 | Components after truncation: 3
WAIC: 918.867
lppd: -350.425 | pWAIC: 109.009
Summary table
parameter mean sd q0.025 q0.500 q0.975 ess
weights[1] 0.417 0.037 0.36 0.415 0.491 33.764
weights[2] 0.355 0.032 0.299 0.35 0.415 59.97
weights[3] 0.229 0.054 0.125 0.238 0.315 33.929
alpha 0.51 0.313 0.121 0.447 1.261 150
location[1] 5.795 2.24 1.026 6.739 7.819 17.997
location[2] 2.387 2.454 0.89 1.109 7.844 30.481
location[3] 2.48 0.68 0.618 2.697 3.034 12.19
scale[1] 0.534 0.46 0.259 0.33 1.7 19.738
scale[2] 1.479 0.66 0.281 1.687 2.457 48.891
scale[3] 1.477 0.567 0.781 1.316 2.78 26.65
Code
<- params (fit_crp)
Posterior mean parameters
$alpha
[1] 0.5099
$w
[1] 0.4167 0.3548 0.2285
$location
[1] 5.795 2.387 2.480
$scale
[1] 0.5344 1.4790 1.4770
MCMC Diagnostics Plots
Code
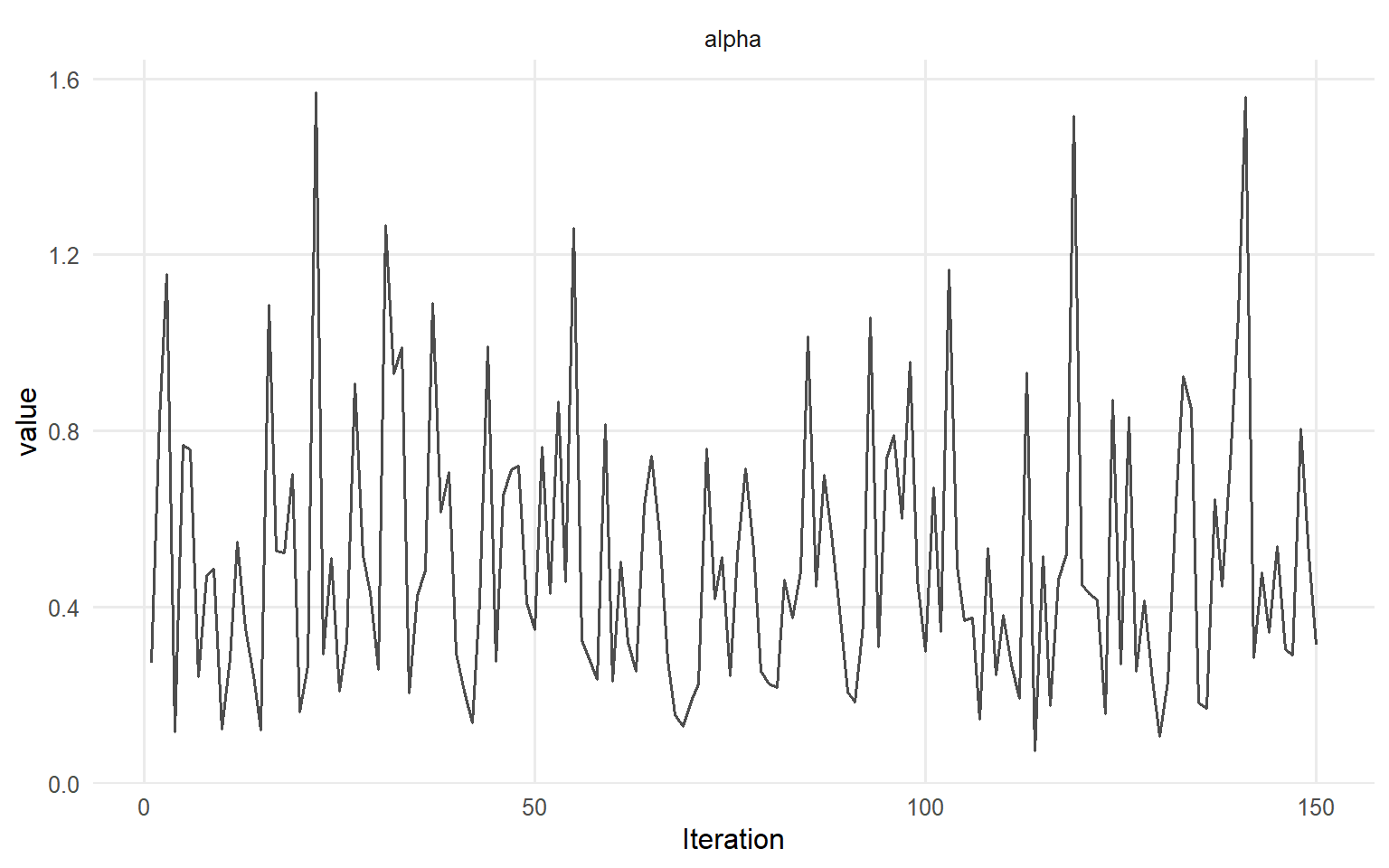
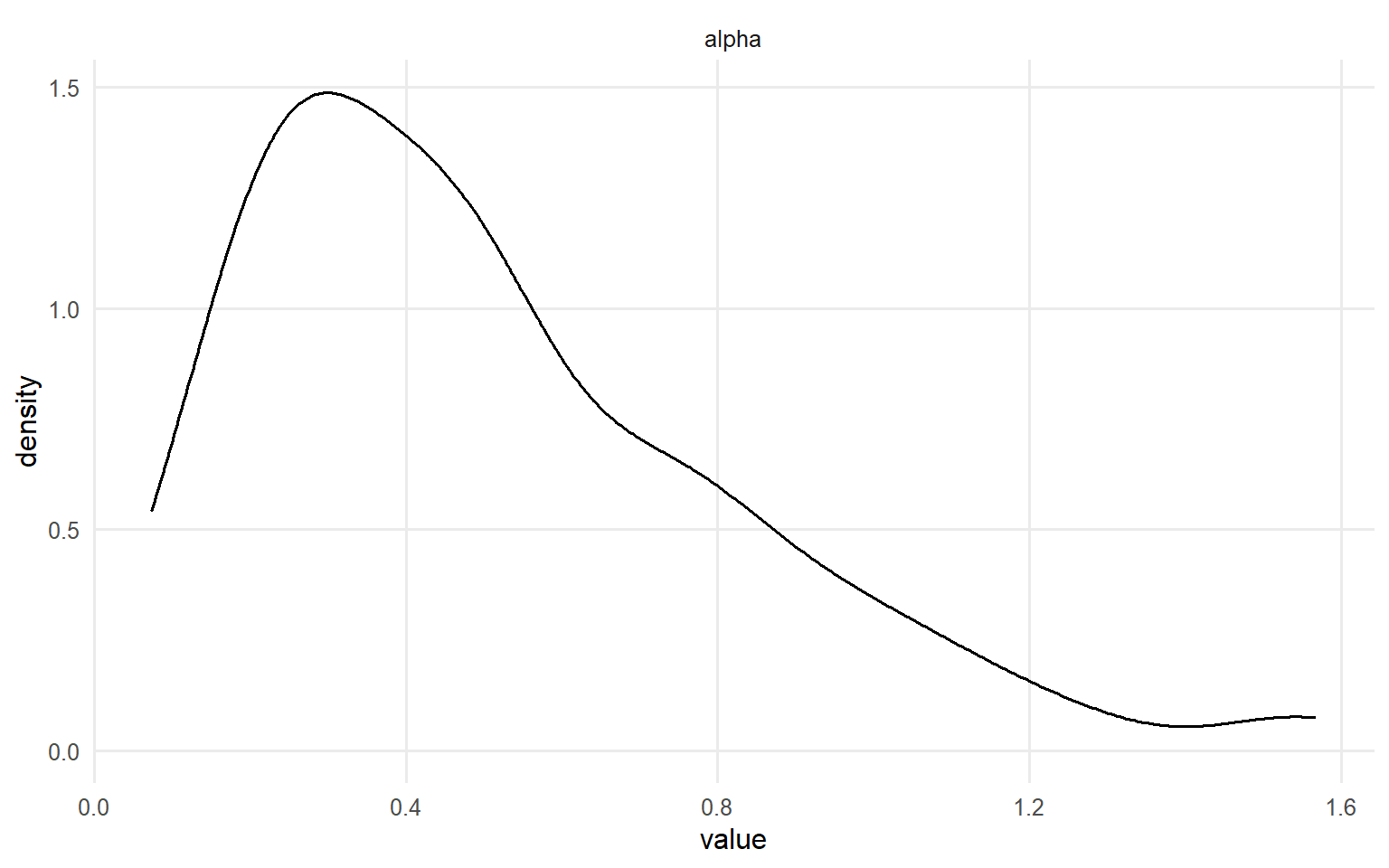
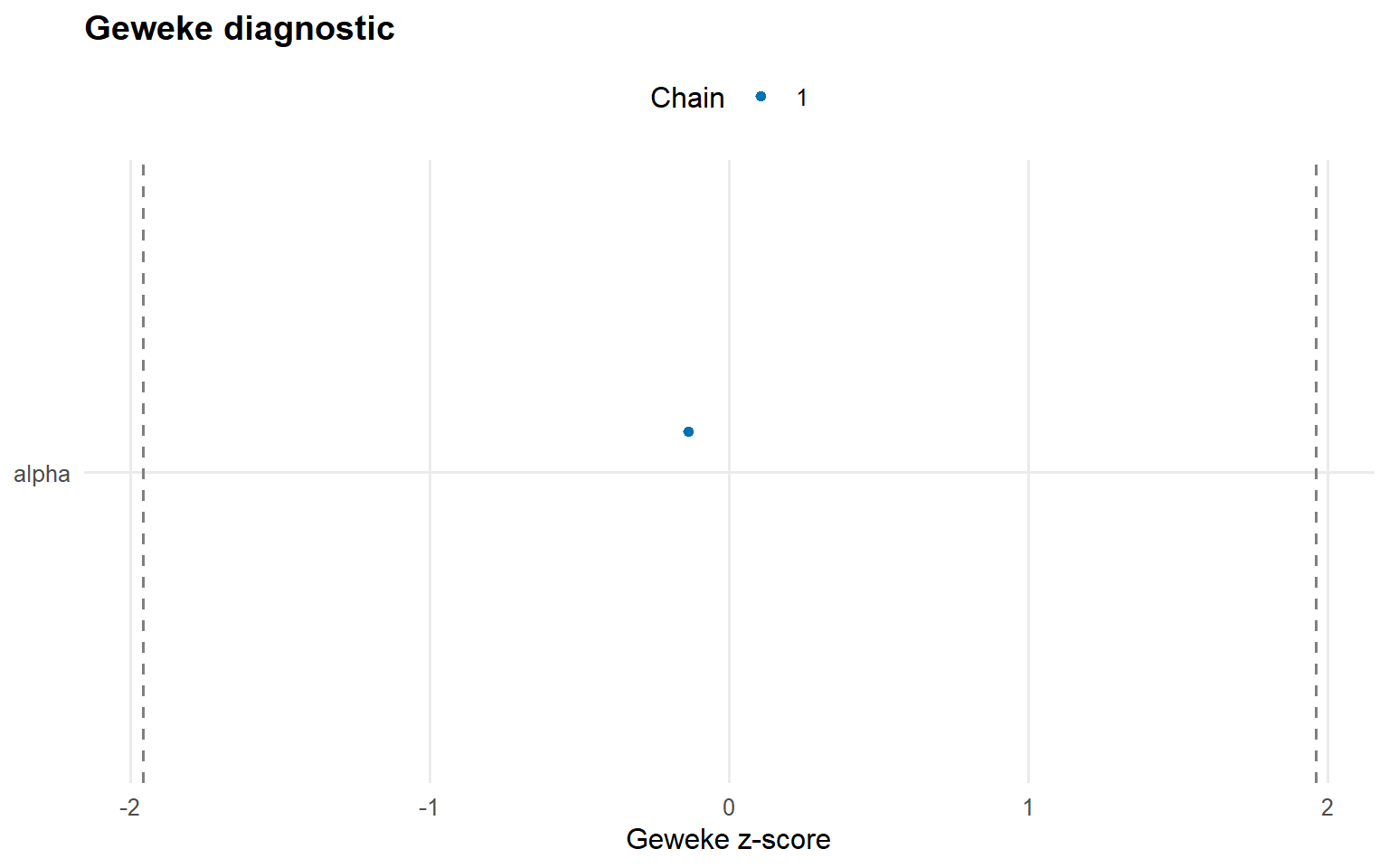
# Trace plots for key parameters plot (fit_crp, params = "alpha" , family = c ("traceplot" , "density" , "geweke" ))
Posterior Predictions
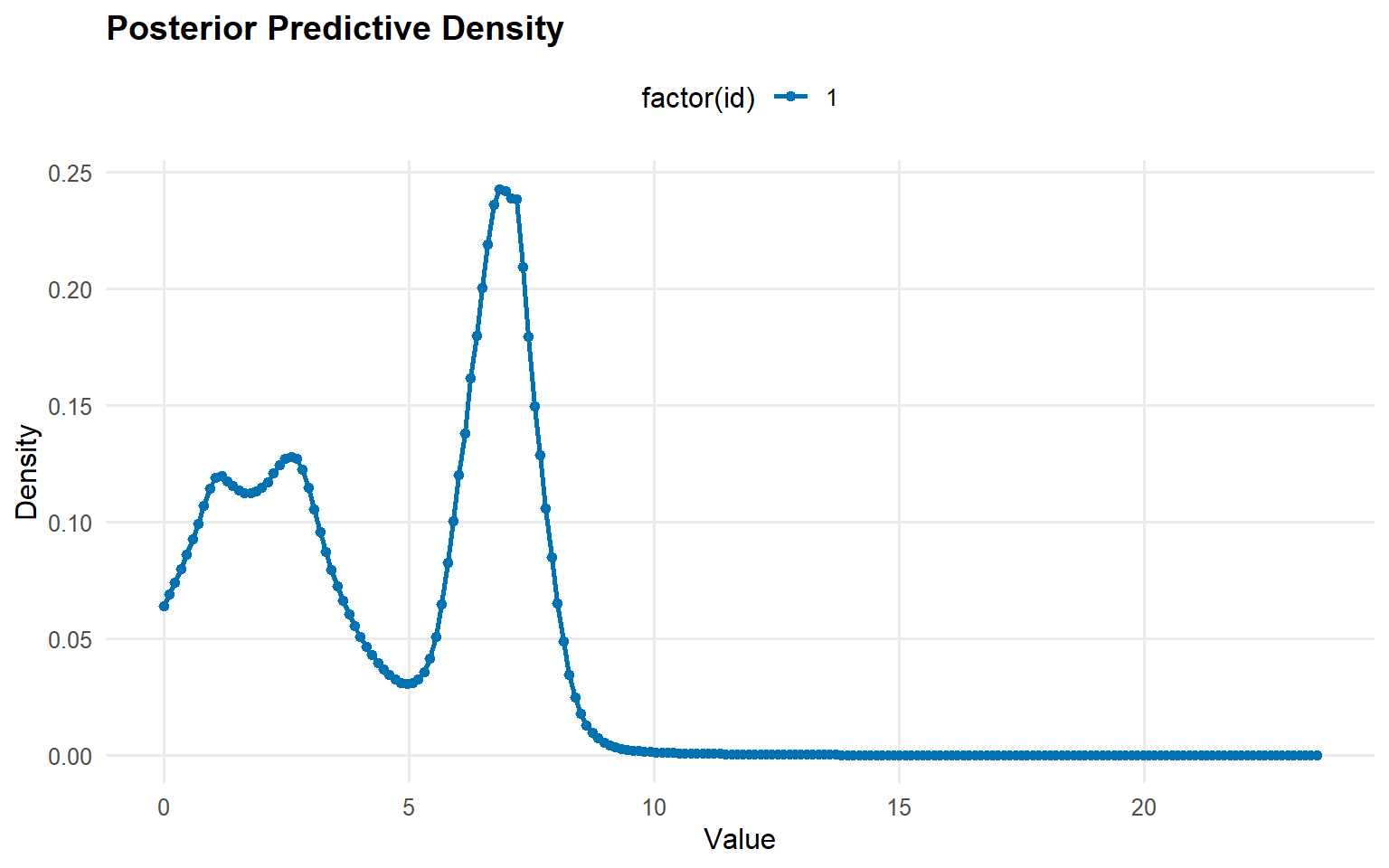
Predictive Density
Code
# Generate prediction grid <- seq (0 , max (y_mixed) * 1.2 , length.out = 200 )# Posterior predictive density <- predict (fit_crp, y = y_grid, type = "density" )# Use S3 plot method plot (pred_density)
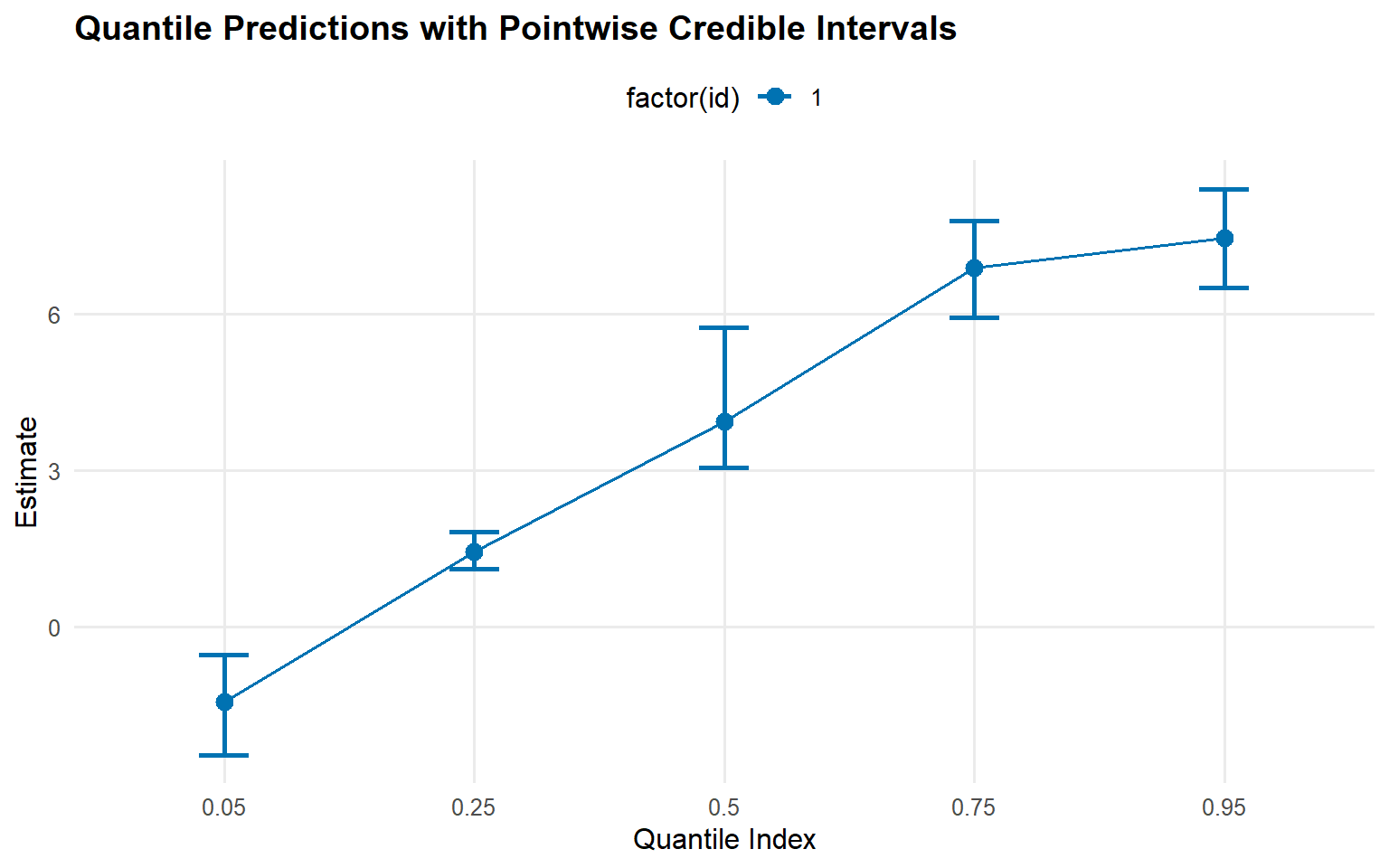
Quantile Predictions
Code
# Posterior predictive quantiles with credible intervals <- predict (fit_crp, type = "quantile" , p = c (0.05 , 0.25 , 0.5 , 0.75 , 0.95 ),interval = "credible" )# Use S3 plot method plot (quantiles_pred)
Varying Truncation Level (components)
Code
# Demonstrate with one value <- bundle (y = y_mixed,kernel = "laplace" ,backend = "crp" ,components = 5 ,mcmc = mcmc<- load_or_fit ("ex01-unconditional-dpm-crp-fit_components" , dpmix (bundle_components))
Code
MixGPD summary | backend: Chinese Restaurant Process | kernel: Laplace Distribution | GPD tail: FALSE | epsilon: 0.025
n = 200 | components = 5
Summary
Initial components: 5 | Components after truncation: 3
WAIC: 873.554
lppd: -291.702 | pWAIC: 145.075
Summary table
parameter mean sd q0.025 q0.500 q0.975 ess
weights[1] 0.351 0.058 0.264 0.335 0.49 24.589
weights[2] 0.268 0.042 0.195 0.27 0.346 31.995
weights[3] 0.201 0.042 0.11 0.2 0.281 9.12
alpha 0.808 0.442 0.183 0.722 1.78 150
location[1] 3.2 2.543 0.803 2.349 7.961 10.681
location[2] 4.566 3.273 0.741 2.824 9.893 11.698
location[3] 4.393 3.496 0.423 2.691 10.3 22.648
scale[1] 1.215 0.686 0.269 1.162 2.352 11.444
scale[2] 1.129 0.793 0.25 1.044 2.791 22.497
scale[3] 1.486 1.113 0.28 1.291 3.987 19.074
For unconditional models, use predict() for posterior summaries; fitted() and residuals() are not supported.
Model Comparison: Different Kernels
Laplace Kernel (Current)
Code
<- bundle (y = y_mixed,kernel = "laplace" ,backend = "crp" ,components = 5 ,mcmc = mcmc<- load_or_fit ("ex01-unconditional-dpm-crp-fit_laplace" , dpmix (bundle_laplace))
Code
MixGPD summary | backend: Chinese Restaurant Process | kernel: Laplace Distribution | GPD tail: FALSE | epsilon: 0.025
n = 200 | components = 5
Summary
Initial components: 5 | Components after truncation: 3
WAIC: 900.436
lppd: -326.205 | pWAIC: 124.013
Summary table
parameter mean sd q0.025 q0.500 q0.975 ess
weights[1] 0.403 0.042 0.307 0.405 0.475 24.123
weights[2] 0.323 0.052 0.229 0.325 0.411 6.766
weights[3] 0.223 0.047 0.119 0.225 0.301 18.724
alpha 0.755 0.431 0.178 0.654 1.752 100.851
location[1] 6.039 2.085 1.15 6.827 7.776 37.299
location[2] 2.794 2.148 0.795 2.305 7.416 37.024
location[3] 1.564 1.098 0.491 1.159 2.745 13.27
scale[1] 0.51 0.421 0.269 0.337 1.773 58.471
scale[2] 1.284 0.585 0.313 1.274 2.392 50.835
scale[3] 2.073 0.819 0.846 2.082 3.777 24.931
Amoroso Kernel (Alternative)
Code
<- bundle (y = y_mixed,kernel = "amoroso" ,backend = "crp" ,components = 5 ,mcmc = mcmc<- load_or_fit ("ex01-unconditional-dpm-crp-fit_amoroso" , dpmix (bundle_amoroso))
Code
MixGPD summary | backend: Chinese Restaurant Process | kernel: Amoroso Distribution | GPD tail: FALSE | epsilon: 0.025
n = 200 | components = 5
Summary
Initial components: 5 | Components after truncation: 2
WAIC: 819.477
lppd: -356.289 | pWAIC: 53.45
Summary table
parameter mean sd q0.025 q0.500 q0.975 ess
weights[1] 0.637 0.08 0.505 0.647 0.781 5.601
weights[2] 0.35 0.082 0.204 0.345 0.485 5.217
alpha 0.549 0.351 0.057 0.487 1.345 72.953
loc[1] -0.002 0.22 -0.112 0.028 0.04 36.674
loc[2] 1.774 1.34 -1.356 2.098 3.74 3.383
scale[1] 2.931 0.52 1.629 3.022 3.749 16.876
scale[2] 1.673 0.644 0.86 1.519 3.202 5.92
shape1[1] 0.647 0.668 0.233 0.603 0.918 52.106
shape1[2] 3.76 1.042 1.923 3.856 5.836 7.17
shape2[1] 2.362 1.173 1.278 1.966 4.822 1.52
shape2[2] 0.977 0.403 0.803 0.945 1.024 58.526
Model Comparison via Predictions
Code
# Compare models using predict() (fitted/residuals not supported for unconditional) <- predict (fit_laplace, type = "mean" , level = 0.9 , interval = "credible" )<- predict (fit_amoroso, type = "mean" , level = 0.9 , interval = "credible" )
Code
# For unconditional models, fitted() is not supported; use predict() for comparisons (see pred_laplace, pred_amoroso above).
Key Takeaways
CRP Backend : Flexible cluster assignment, ideal for unknown mixture complexitycomponents Parameter : A higher components cap allows more represented mixture components but increases computationKernel Choice : Laplace and Amoroso illustrate two different bulk-kernel choices for the same unconditional CRP workflowDiagnostics : Check convergence (Rhat, ESS) and posterior predictive fitNext : Compare with Stick-Breaking (SB) backend in ex02
Prereqs
Required packages and data for this page are listed in the setup chunks above.
Outputs
This page renders model fits, diagnostics, and summary artifacts generated by package APIs.
Interpretation
Canonical concept page: Model Umbrella
Treat this page as an application/example view and use the canonical page for core definitions.
Next
Continue to the linked canonical concept page, then return for implementation-specific details.