$fit
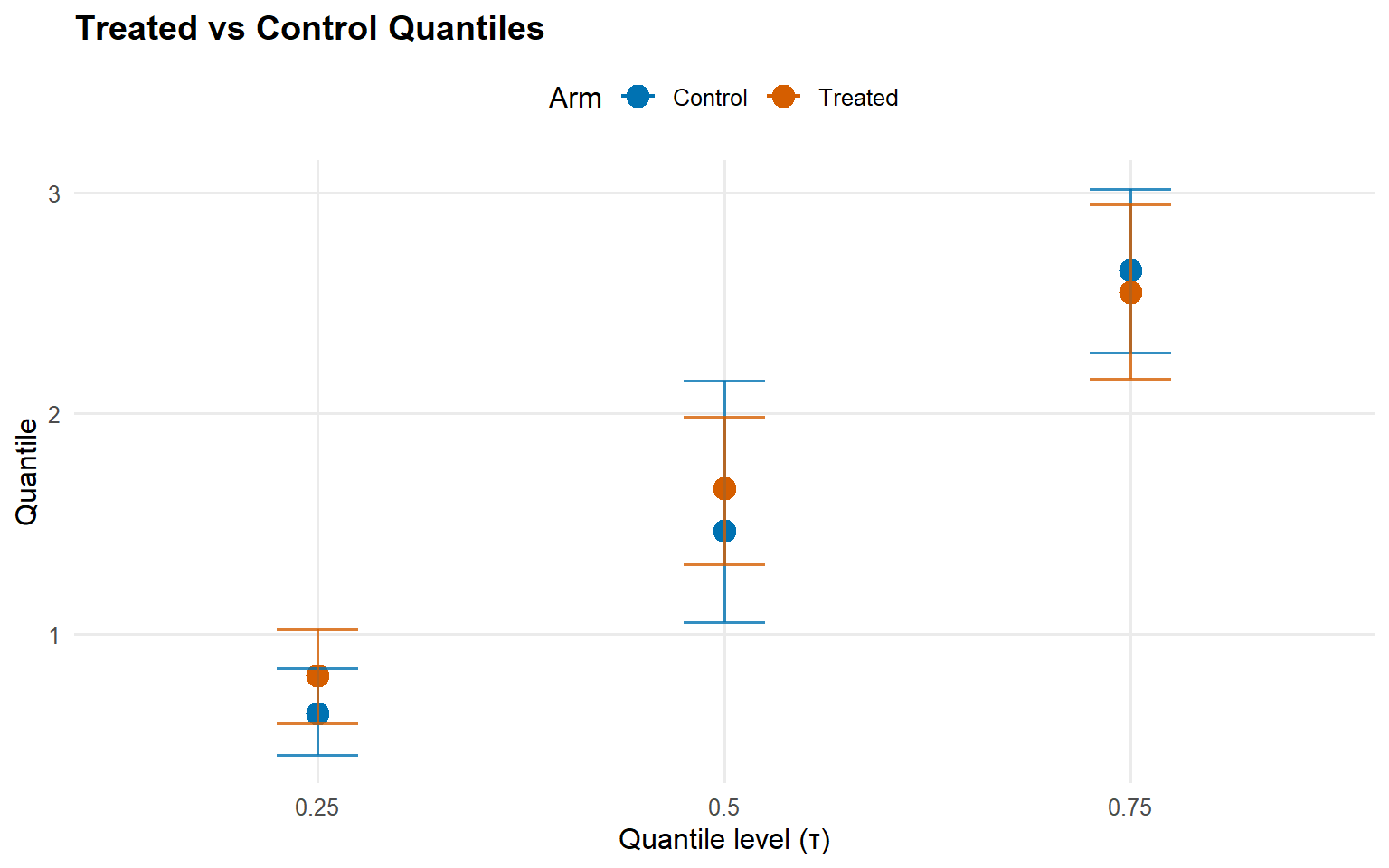
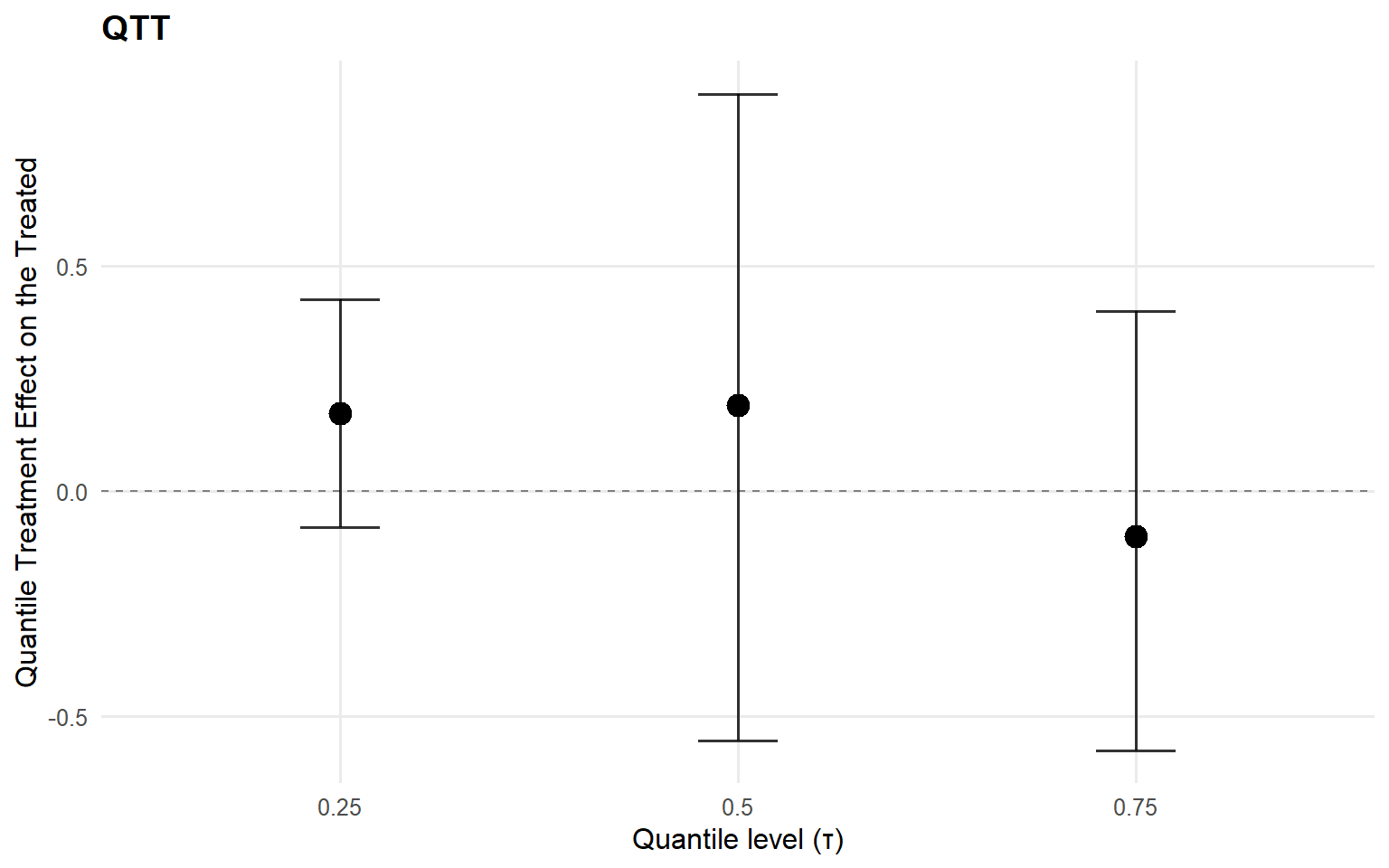
index estimate lower upper
1 0.25 0.17270432 -0.07889603 0.4250001
2 0.50 0.19139716 -0.55257196 0.8816509
3 0.75 -0.09985004 -0.57517407 0.3992333
$fit_df
index estimate lower upper
1 0.25 0.17270432 -0.07889603 0.4250001
2 0.50 0.19139716 -0.55257196 0.8816509
3 0.75 -0.09985004 -0.57517407 0.3992333
$lower
[1] -0.07889603 -0.55257196 -0.57517407
$upper
[1] 0.4250001 0.8816509 0.3992333
$qte
$qte$fit
index estimate lower upper
1 0.25 0.17270432 -0.07889603 0.4250001
2 0.50 0.19139716 -0.55257196 0.8816509
3 0.75 -0.09985004 -0.57517407 0.3992333
$qte$draws
, , 1
[,1]
[1,] 0.1287629791
[2,] -0.1332456252
[3,] 0.0303336012
[4,] -0.0184126278
[5,] -0.0193407294
[6,] -0.0343073323
[7,] 0.0086856266
[8,] 0.2455021374
[9,] 0.1937363934
[10,] 0.2587364507
[11,] 0.1832300251
[12,] 0.2060050759
[13,] 0.2342316412
[14,] 0.2240440824
[15,] 0.2610187324
[16,] 0.2865283574
[17,] 0.3270466410
[18,] 0.3523919852
[19,] 0.3738278154
[20,] 0.2559115494
[21,] 0.1774994291
[22,] 0.1811404731
[23,] 0.0039228043
[24,] 0.2912818762
[25,] 0.0946364319
[26,] 0.1387677086
[27,] 0.0426923925
[28,] -0.0618880886
[29,] -0.0942841633
[30,] 0.0393527873
[31,] 0.1506993825
[32,] 0.1773688647
[33,] 0.2827492349
[34,] 0.3048613636
[35,] 0.1878716395
[36,] 0.0001280459
[37,] 0.0259096352
[38,] 0.0328451108
[39,] -0.0611695272
[40,] 0.1811980530
[41,] 0.3195630237
[42,] 0.1904094799
[43,] -0.0085487156
[44,] 0.3017957765
[45,] 0.3263986781
[46,] 0.0095750997
[47,] -0.0152571492
[48,] 0.3283926775
[49,] 0.4406240698
[50,] 0.3539425185
[51,] 0.3674717267
[52,] 0.1909473598
[53,] 0.4164770600
[54,] 0.3959569032
[55,] 0.3654314617
[56,] 0.1348144967
[57,] 0.1495632509
[58,] -0.0027853762
[59,] 0.4327113775
[60,] 0.2045050004
, , 2
[,1]
[1,] -0.05661537
[2,] -0.38296149
[3,] 0.18046696
[4,] 0.07126461
[5,] 0.02666179
[6,] -0.37718641
[7,] -0.35336540
[8,] 0.59679425
[9,] 0.38786398
[10,] 0.59178232
[11,] 0.55829702
[12,] 0.48718922
[13,] 0.53481636
[14,] 0.51472751
[15,] 0.64403570
[16,] 0.48734074
[17,] 0.52672166
[18,] 0.38109749
[19,] 0.53765255
[20,] 0.31192325
[21,] 0.03260523
[22,] 0.16571383
[23,] 0.35600833
[24,] 0.25736513
[25,] 0.49431465
[26,] 0.27350717
[27,] -0.09735138
[28,] -0.12642415
[29,] -0.59498826
[30,] -0.60338652
[31,] -0.24120519
[32,] 0.11931928
[33,] 0.20452304
[34,] 0.29077214
[35,] -0.17550763
[36,] -0.16904331
[37,] -0.26575729
[38,] 0.02990274
[39,] -0.44058177
[40,] -0.21021453
[41,] -0.08347455
[42,] 0.02607906
[43,] -0.50569079
[44,] -0.01409299
[45,] 0.36588857
[46,] 0.09086234
[47,] -0.08357356
[48,] 0.38926359
[49,] 0.51426764
[50,] 0.50679719
[51,] 0.75059088
[52,] 0.03744930
[53,] 0.94098923
[54,] 0.88477862
[55,] 0.68018352
[56,] 0.24615700
[57,] 0.16264838
[58,] -0.04916649
[59,] 0.87819404
[60,] 0.77760011
, , 3
[,1]
[1,] -0.4156390441
[2,] -0.3979544016
[3,] 0.1864923993
[4,] -0.1158334980
[5,] -0.2543406067
[6,] -0.1533826190
[7,] -0.0835053305
[8,] -0.0238024246
[9,] -0.1372273029
[10,] -0.0845652455
[11,] 0.2473862975
[12,] -0.1541927597
[13,] 0.0642517269
[14,] 0.0394808504
[15,] -0.2203044377
[16,] -0.5503861385
[17,] 0.0821251880
[18,] -0.0671572545
[19,] -0.2722840432
[20,] 0.0239177199
[21,] -0.3788684946
[22,] -0.1698056531
[23,] -0.0621853727
[24,] -0.1756100103
[25,] -0.1738214783
[26,] -0.1889882676
[27,] -0.0759817333
[28,] -0.0098277802
[29,] -0.5976012457
[30,] -0.5044798468
[31,] -0.3197143548
[32,] 0.0435991303
[33,] -0.0102651298
[34,] -0.0973713251
[35,] -0.0775018136
[36,] -0.1905600132
[37,] -0.1390878377
[38,] -0.2466176731
[39,] -0.4358974768
[40,] -0.4885181643
[41,] 0.0612279426
[42,] -0.0068211441
[43,] -0.7174678578
[44,] -0.1307587403
[45,] 0.1632387303
[46,] -0.0002681153
[47,] -0.3939357136
[48,] 0.2191277678
[49,] 0.3303920445
[50,] 0.0363143711
[51,] 0.2520918968
[52,] 0.2350462384
[53,] 0.0403480683
[54,] 0.3874045624
[55,] 0.0956619237
[56,] -0.1702222530
[57,] -0.1431654327
[58,] -0.5296427806
[59,] 0.4099355421
[60,] 0.4565157643
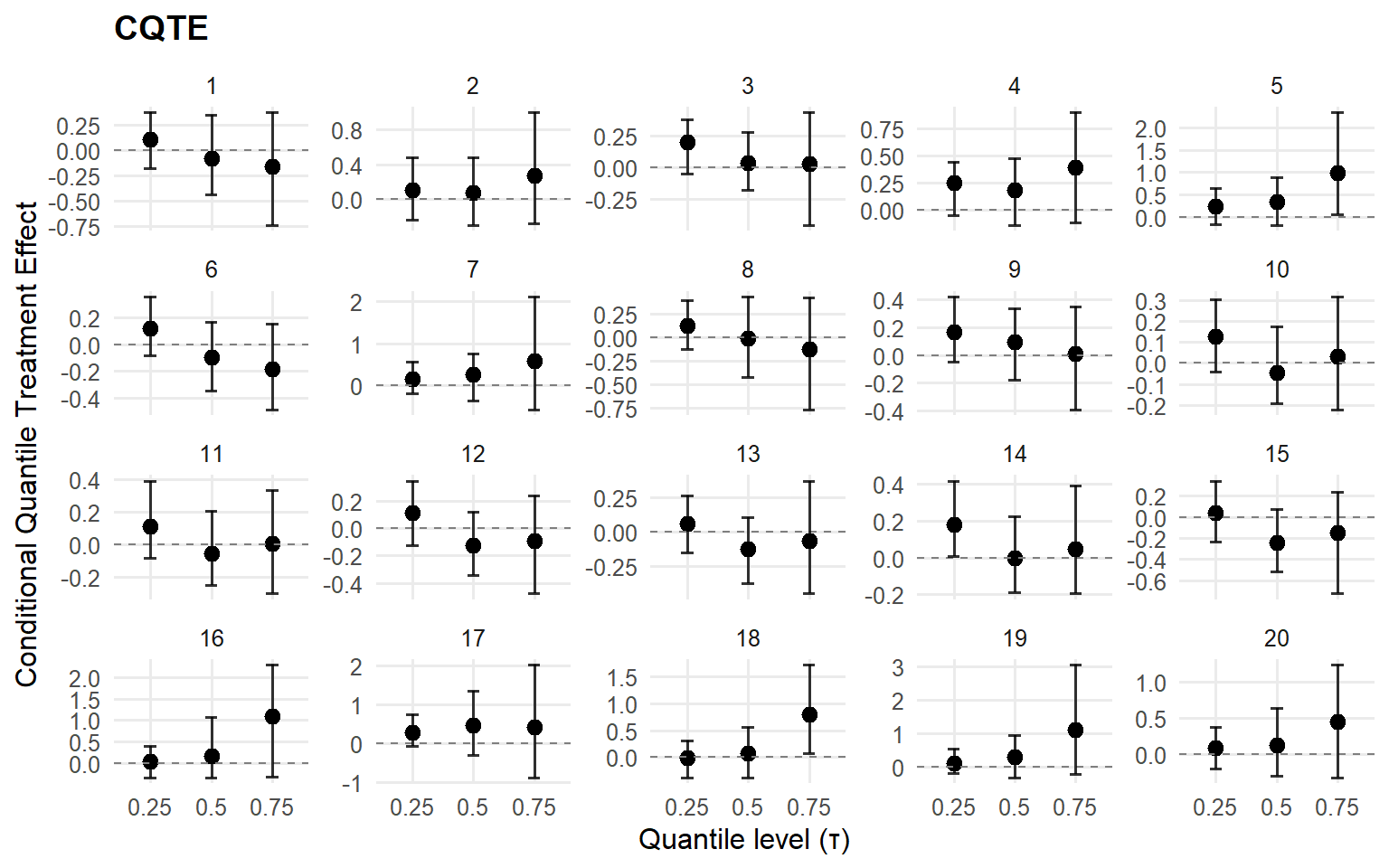
$probs
[1] 0.25 0.50 0.75