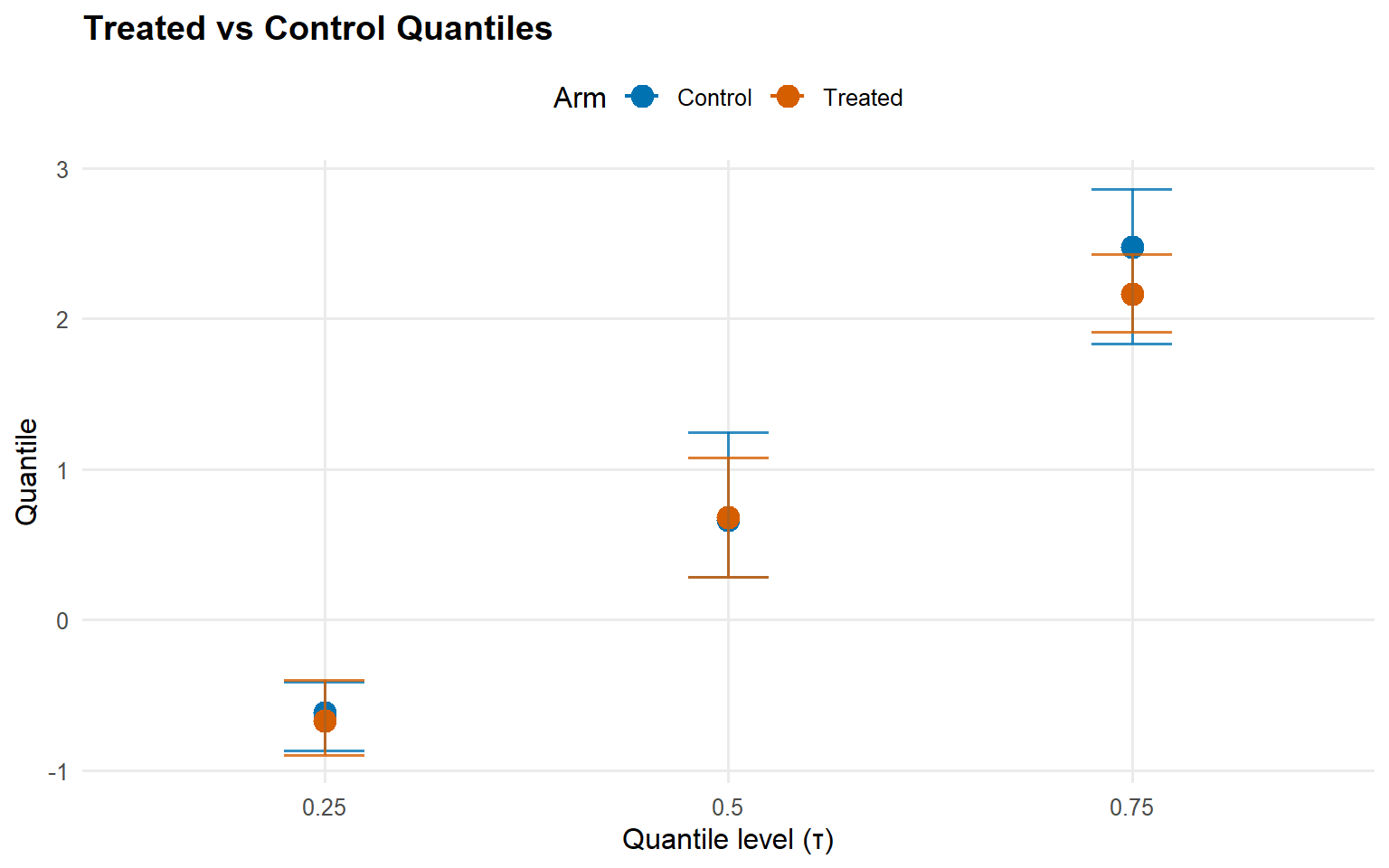
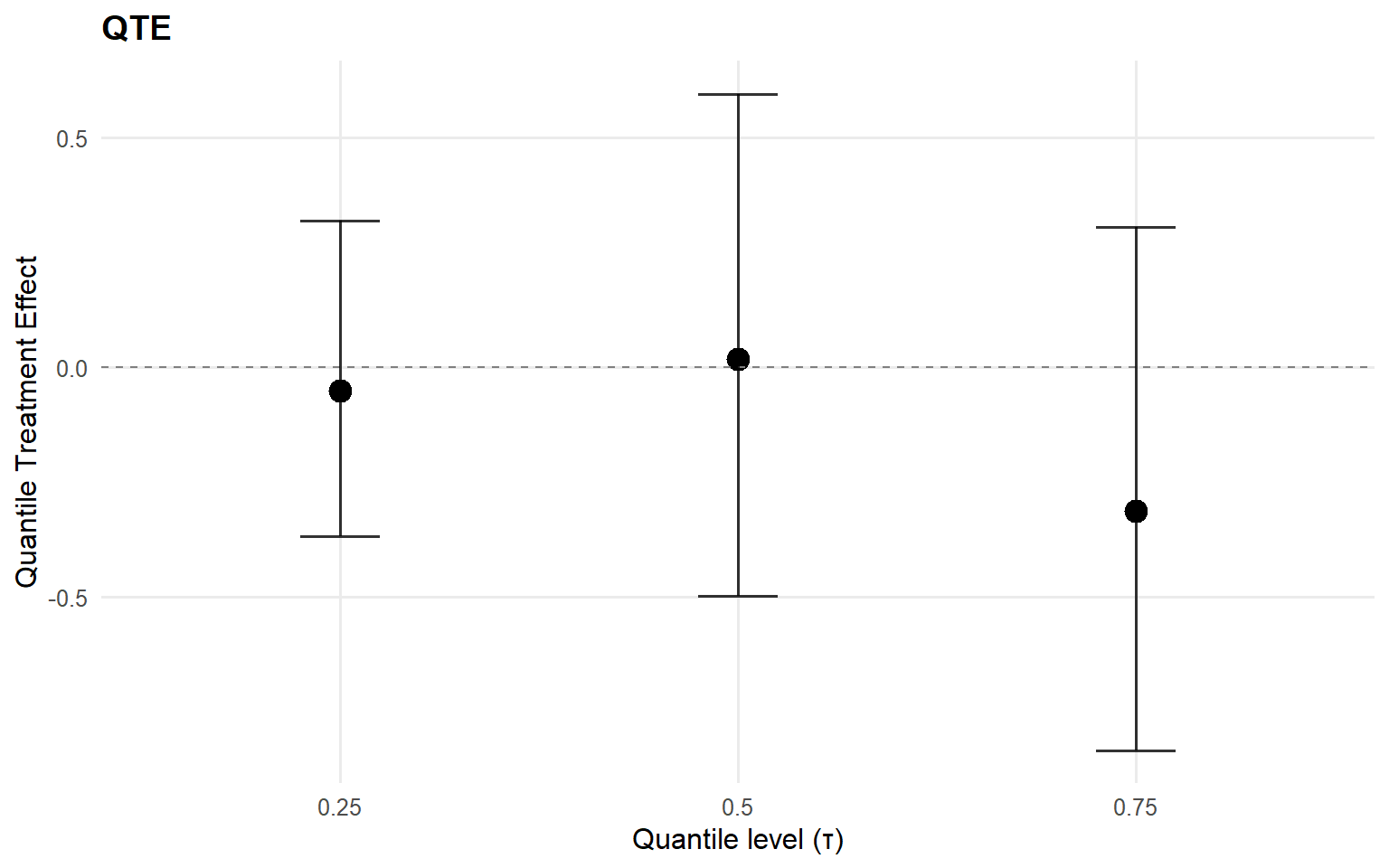
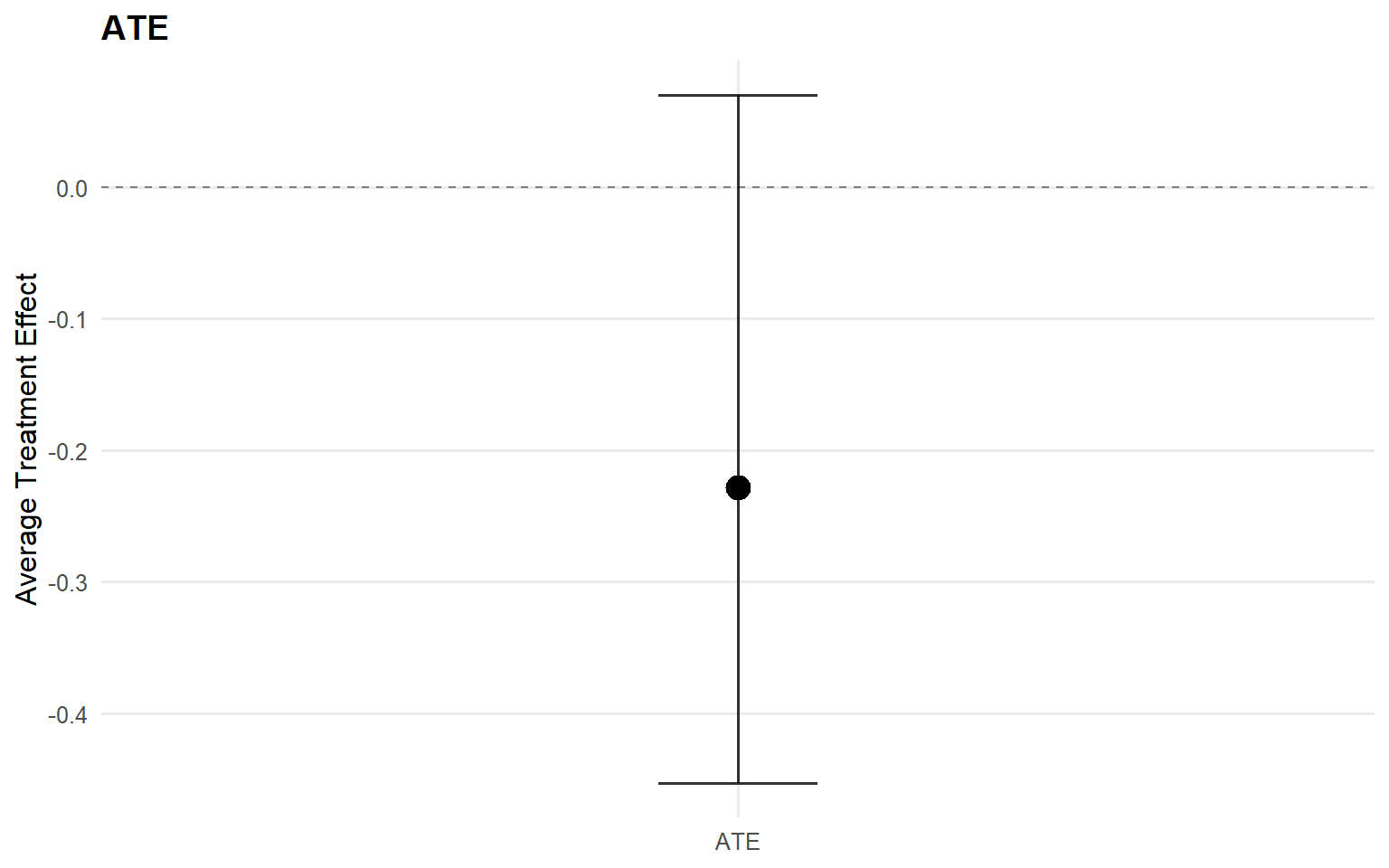$fit
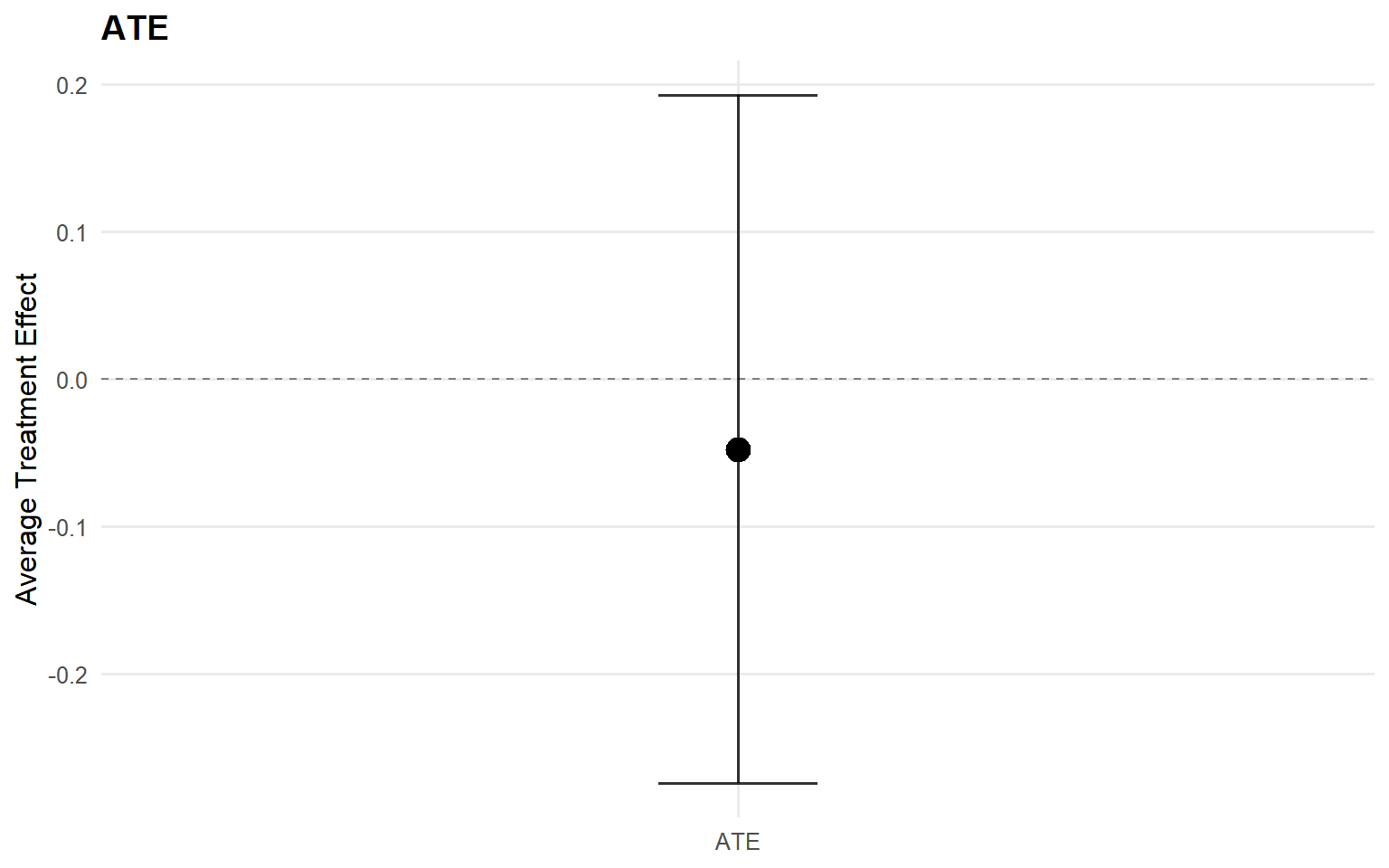
index estimate lower upper
1 0.25 -0.03453822 -0.4578704 0.3000317
2 0.50 -0.06214089 -0.4803350 0.3144029
3 0.75 -0.11304528 -0.7049802 0.4147881
$fit_df
index estimate lower upper
1 0.25 -0.03453822 -0.4578704 0.3000317
2 0.50 -0.06214089 -0.4803350 0.3144029
3 0.75 -0.11304528 -0.7049802 0.4147881
$lower
[1] -0.4578704 -0.4803350 -0.7049802
$upper
[1] 0.3000317 0.3144029 0.4147881
$qte
$qte$fit
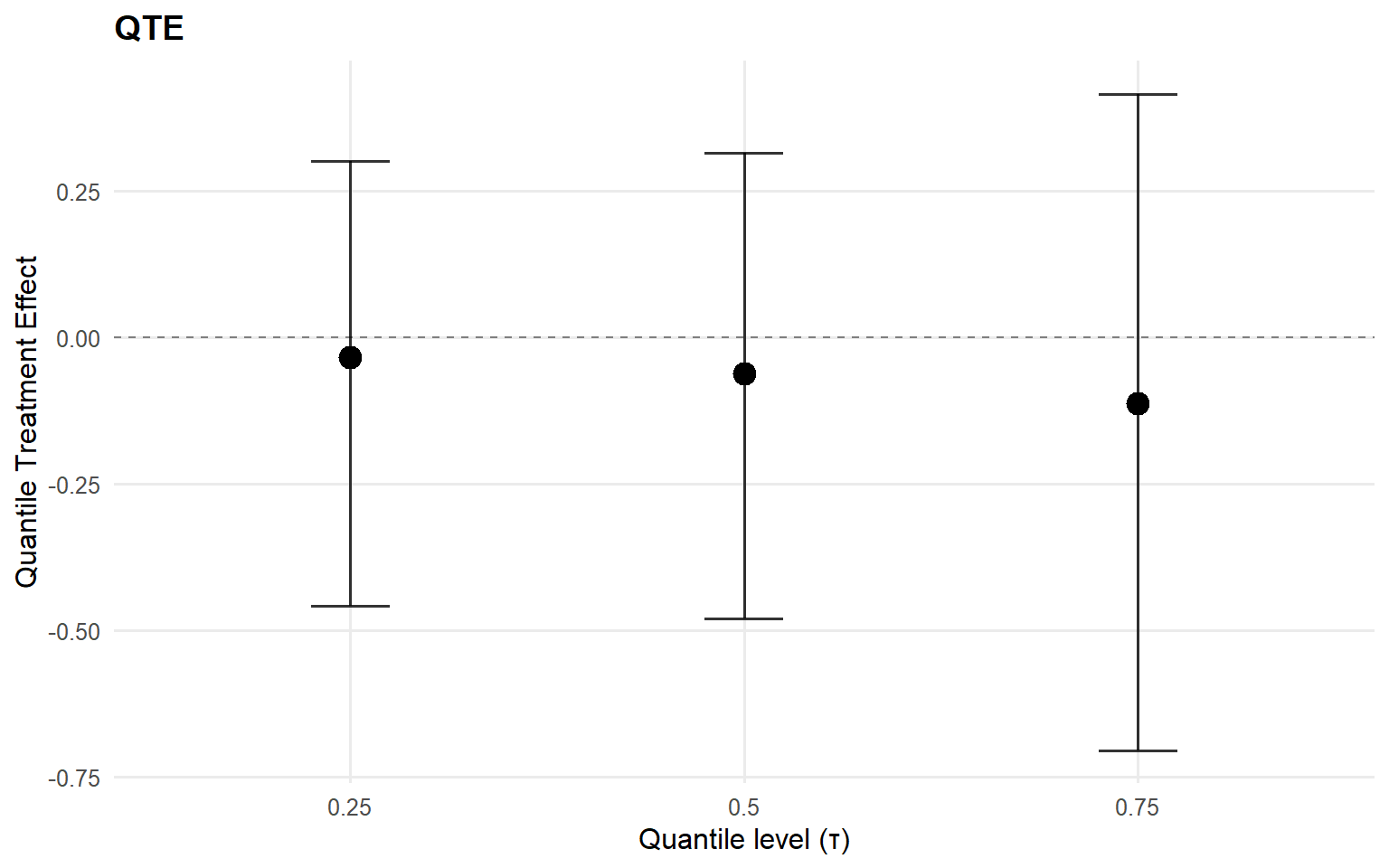
index estimate lower upper
1 0.25 -0.03453822 -0.4578704 0.3000317
2 0.50 -0.06214089 -0.4803350 0.3144029
3 0.75 -0.11304528 -0.7049802 0.4147881
$qte$draws
, , 1
[,1]
[1,] 0.043709335
[2,] 0.131132709
[3,] 0.223139040
[4,] 0.117772385
[5,] -0.356922199
[6,] 0.131853557
[7,] -0.071811459
[8,] -0.268375775
[9,] -0.189619370
[10,] 0.020175368
[11,] -0.445303737
[12,] -0.279374656
[13,] -0.042429918
[14,] -0.233787449
[15,] 0.230592156
[16,] 0.128408519
[17,] -0.254115497
[18,] 0.083323483
[19,] -0.067695918
[20,] -0.284575899
[21,] -0.207634706
[22,] 0.135321424
[23,] 0.152477120
[24,] 0.011546942
[25,] -0.130472065
[26,] -0.241793239
[27,] -0.015708168
[28,] -0.265357590
[29,] -0.112670495
[30,] -0.433802411
[31,] -0.351310280
[32,] 0.053355858
[33,] 0.444951931
[34,] -0.117841403
[35,] 0.130488639
[36,] -0.118774557
[37,] 0.038011868
[38,] 0.150523301
[39,] -0.123756943
[40,] -0.108957156
[41,] 0.030067292
[42,] 0.172131137
[43,] -0.104723073
[44,] -0.031876979
[45,] -0.040161736
[46,] -0.234373450
[47,] -0.296631797
[48,] -0.333291968
[49,] -0.716884129
[50,] -0.056477246
[51,] 0.016758376
[52,] -0.242433862
[53,] -0.020528307
[54,] -0.354413143
[55,] -0.104246822
[56,] -0.109817755
[57,] -0.461518842
[58,] -0.051546640
[59,] -0.061326124
[60,] -0.025922822
[61,] -0.305430651
[62,] 0.157299935
[63,] 0.213135433
[64,] 0.261618424
[65,] -0.235931321
[66,] 0.189187374
[67,] 0.065764865
[68,] 0.322942179
[69,] 0.236925413
[70,] 0.262666267
[71,] 0.227627831
[72,] 0.072452963
[73,] -0.035905719
[74,] 0.284279289
[75,] 0.198222762
[76,] 0.264292589
[77,] 0.117708391
[78,] 0.178460277
[79,] 0.194520420
[80,] 0.304605020
[81,] 0.158399487
[82,] -0.158591338
[83,] -0.493598675
[84,] -0.309140975
[85,] -0.005371278
[86,] -0.222657916
[87,] 0.263461308
[88,] -0.004638298
[89,] 0.142520254
[90,] 0.069260893
, , 2
[,1]
[1,] -0.065371036
[2,] 0.122850097
[3,] 0.138547360
[4,] -0.027724289
[5,] 0.207264000
[6,] 0.175053598
[7,] 0.027986521
[8,] -0.016130181
[9,] -0.076791362
[10,] 0.205566431
[11,] -0.119453679
[12,] 0.008763608
[13,] 0.130124422
[14,] 0.140328222
[15,] 0.401856484
[16,] 0.008393922
[17,] 0.132916654
[18,] 0.177798716
[19,] -0.005123042
[20,] 0.005964241
[21,] 0.012415567
[22,] 0.195577178
[23,] 0.323908706
[24,] 0.074032983
[25,] -0.240974434
[26,] -0.020079897
[27,] 0.045370100
[28,] -0.358455773
[29,] -0.127090303
[30,] -0.264249075
[31,] -0.291660631
[32,] 0.024944847
[33,] 0.263351387
[34,] -0.062925761
[35,] 0.263648953
[36,] 0.029644997
[37,] 0.281660675
[38,] 0.074463447
[39,] -0.267513576
[40,] -0.115188857
[41,] -0.099831722
[42,] 0.177389597
[43,] -0.292281724
[44,] -0.141140938
[45,] 0.052752714
[46,] -0.128914176
[47,] -0.322449261
[48,] -0.379400372
[49,] -0.484852105
[50,] -0.146664139
[51,] -0.012908775
[52,] -0.194473472
[53,] -0.247253347
[54,] -0.344647872
[55,] -0.332726347
[56,] -0.172702873
[57,] -0.541544732
[58,] -0.275711458
[59,] -0.184467309
[60,] -0.206013791
[61,] -0.511302527
[62,] -0.032871555
[63,] -0.083448788
[64,] 0.005566095
[65,] -0.144513366
[66,] -0.034712794
[67,] -0.188705044
[68,] -0.055884054
[69,] -0.034566013
[70,] -0.164218293
[71,] 0.020301461
[72,] 0.006626609
[73,] -0.221535353
[74,] -0.068373365
[75,] -0.253423434
[76,] -0.301155738
[77,] -0.196817249
[78,] -0.088412120
[79,] -0.044244479
[80,] -0.091236139
[81,] -0.045118855
[82,] -0.069462382
[83,] -0.464776006
[84,] -0.162245798
[85,] -0.061906723
[86,] -0.083088362
[87,] 0.379617788
[88,] 0.100597913
[89,] 0.202037154
[90,] -0.041267986
, , 3
[,1]
[1,] 0.125069433
[2,] -0.079291958
[3,] 0.061382307
[4,] -0.117071563
[5,] 0.194923252
[6,] 0.204081390
[7,] 0.090941193
[8,] -0.047887378
[9,] 0.168778281
[10,] 0.036549058
[11,] -0.114063686
[12,] 0.053409353
[13,] 0.167271594
[14,] 0.320082267
[15,] 0.563068683
[16,] -0.132445704
[17,] 0.209597579
[18,] 0.278072982
[19,] 0.262074233
[20,] 0.219877820
[21,] -0.023930233
[22,] 0.431074409
[23,] 0.153968020
[24,] 0.232282882
[25,] -0.252403904
[26,] -0.040640694
[27,] -0.040006720
[28,] -0.464171648
[29,] -0.163507798
[30,] -0.197037966
[31,] -0.290749321
[32,] -0.105609197
[33,] 0.079741373
[34,] 0.004690825
[35,] 0.107356497
[36,] -0.140186687
[37,] -0.142708648
[38,] 0.036974408
[39,] -0.260562471
[40,] -0.350733344
[41,] -0.270009885
[42,] 0.060835676
[43,] -0.425195834
[44,] -0.283117511
[45,] -0.077086537
[46,] -0.336163228
[47,] -0.315834660
[48,] -0.218022803
[49,] -0.178871011
[50,] 0.194201492
[51,] 0.068150430
[52,] 0.062875725
[53,] -0.359905515
[54,] -0.480115959
[55,] -0.450091094
[56,] -0.201264587
[57,] -0.759053712
[58,] -0.316013618
[59,] -0.302979285
[60,] -0.566736815
[61,] -0.712534115
[62,] -0.239377109
[63,] -0.192689151
[64,] -0.001776688
[65,] -0.296017567
[66,] -0.496354453
[67,] -0.435859943
[68,] -0.153837323
[69,] -0.168944004
[70,] -0.416789638
[71,] -0.240833829
[72,] -0.121197377
[73,] -0.314002097
[74,] -0.678961025
[75,] -0.657219819
[76,] -0.758419706
[77,] -0.371974035
[78,] -0.107933092
[79,] -0.159657643
[80,] 0.022188070
[81,] 0.086352292
[82,] -0.041552097
[83,] -0.284751662
[84,] -0.068537057
[85,] -0.025491238
[86,] 0.358690819
[87,] 0.438239729
[88,] 0.034547195
[89,] 0.119477967
[90,] -0.172721132
$probs
[1] 0.25 0.50 0.75