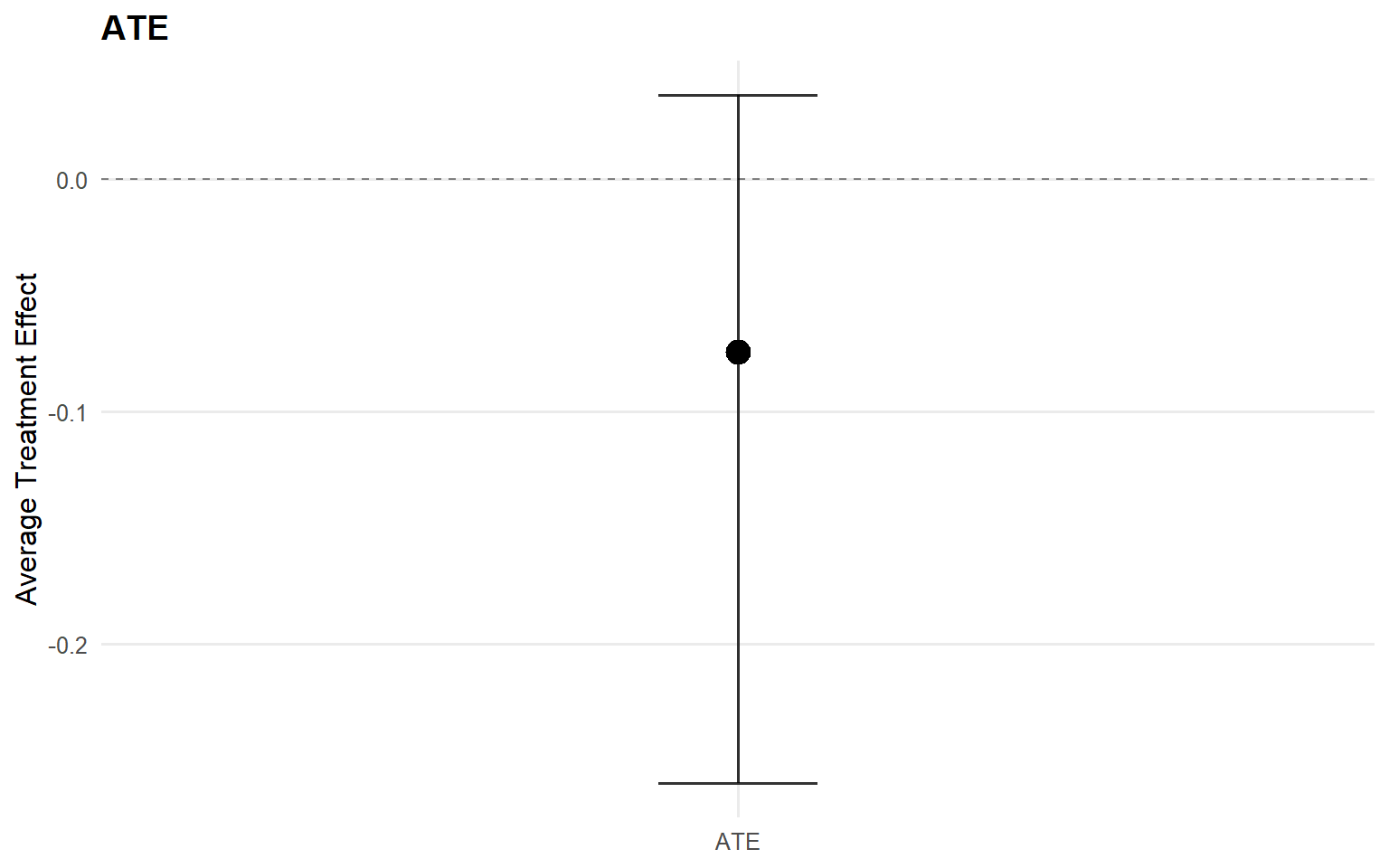Website workflow note. This page reflects the current exported API and recommended wrapper-first usage. Last updated: 2026-02-19 .
For the full package narrative, see the main package vignettes (basic, unconditional, conditional, and causal).
Causal Inference: Same Backend (SB) - Laplace Kernel
This vignette fits two SB-based causal models using the same kernel (Laplace):
Model A: bulk-only outcome models (GPD = FALSE)
Model B: GPD-augmented outcome models (GPD = TRUE)
What you’ll learn
How to hold backend + kernel fixed and isolate the impact of tail augmentation (GPD = FALSE vs TRUE).
How tail augmentation can change conclusions about upper-quantile behavior even when mean effects look similar.
How to run the standard causal post-fit workflow for multiple estimands (mean, quantiles, conditional effects when (X) is present).
When to use this template
You want a clean sensitivity check focused on tail modeling rather than backend/kernel changes.
You want predictable runtime and an explicit complexity knob (components) inside a causal workflow.
Data Setup
Code
data ("causal_alt_real500_p4_k2" )<- causal_alt_real500_p4_k2$ y<- causal_alt_real500_p4_k2$ A<- as.matrix (causal_alt_real500_p4_k2$ X)<- tibble (statistic = c ("N" , "Mean" , "SD" , "Min" , "Max" ),value = c (length (y), mean (y), sd (y), min (y), max (y))
# A tibble: 5 × 2
statistic value
<chr> <dbl>
1 N 500
2 Mean 0.274
3 SD 1.76
4 Min -8.09
5 Max 5.27
Code
<- X[seq_len (if (isTRUE (FAST)) 20 L else 40 L), , drop = FALSE ]<- y[seq_len (if (isTRUE (FAST)) 20 L else 40 L)]<- as.numeric (stats:: quantile (y, 0.8 , names = FALSE ))
Code
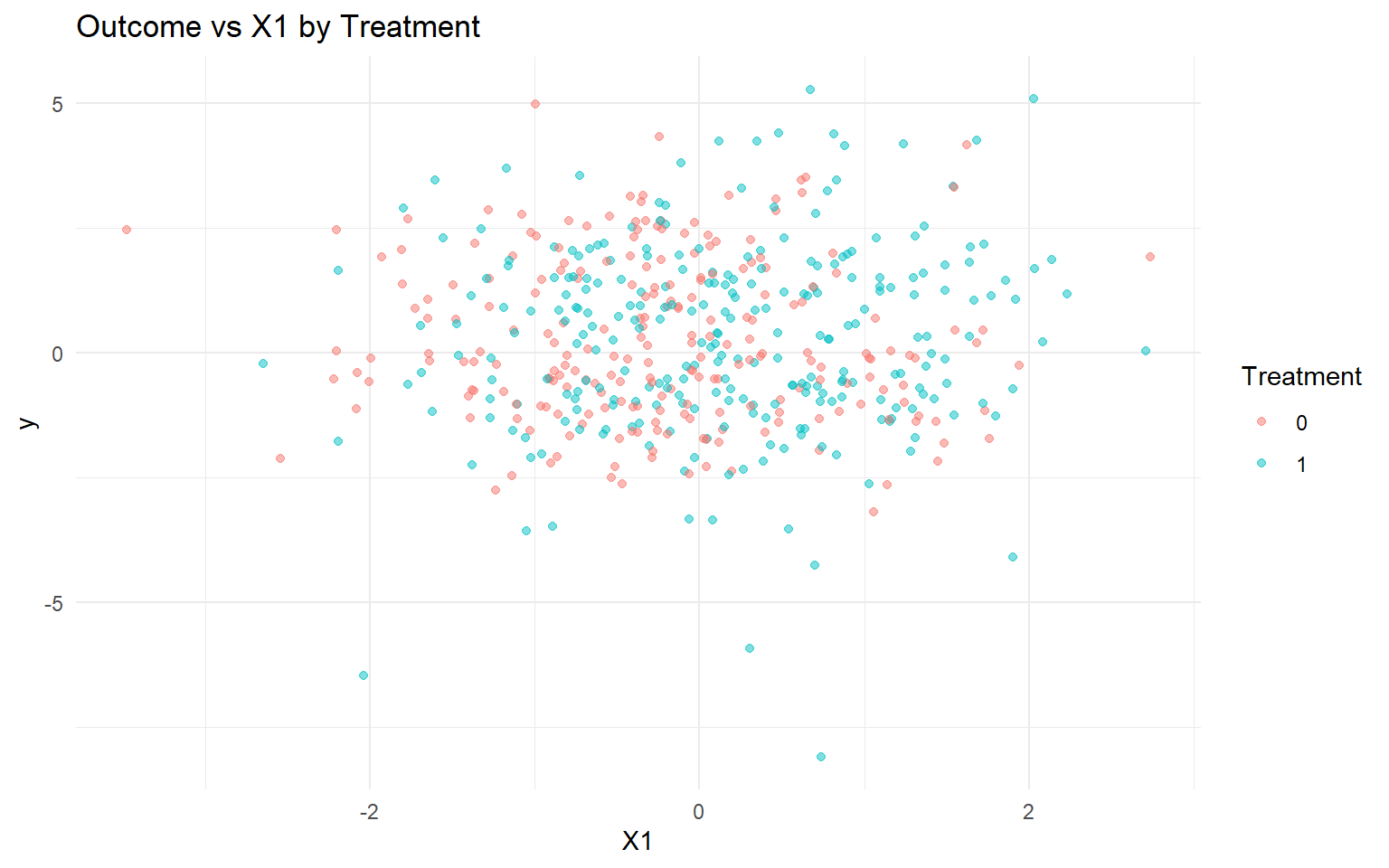
<- data.frame (y = y, A = as.factor (A), x1 = X[, 1 ], x2 = X[, 2 ])<- ggplot (df_causal, aes (x = x1, y = y, color = A)) + geom_point (alpha = 0.5 ) + labs (title = "Outcome vs X1 by Treatment" , x = "X1" , y = "y" , color = "Treatment" ) + theme_minimal ()
Model A: SB Bulk-only (Laplace)
Code
<- bundle (y = y,treat = A,X = X,kernel = "laplace" ,backend = "sb" ,PS = "logit" ,GPD = FALSE ,components = components_ex12,mcmc_outcome = bulk_mcmc_ex12,mcmc_ps = mcmc_ps_bulk_ex12
CausalMixGPD causal bundle
PS model: Bayesian logit (A | X)
Field Treated Control
Backend Stick-Breaking Process Stick-Breaking Process
Kernel laplace laplace
Components 3 3
GPD tail FALSE FALSE
Epsilon 0.025 0.025
Outcome PS included: TRUE
n (control) = 232 | n (treated) = 268
Code
CausalMixGPD causal bundle summary
CausalMixGPD causal bundle
PS model: Bayesian logit (A | X)
Field Treated Control
Backend Stick-Breaking Process Stick-Breaking Process
Kernel laplace laplace
Components 3 3
GPD tail FALSE FALSE
Epsilon 0.025 0.025
Outcome PS included: TRUE
n (control) = 232 | n (treated) = 268
Code
<- load_or_fit (quiet_mcmc (dpmix.causal (bundle_sb_bulk, mcmc = causal_run_mcmc_bulk))summary (fit_sb_bulk)
-- PS fit --
CausalMixGPD PS fit summary
model: logit
n = 500 | predictors = 5
Monitors: beta
Summary table
parameter mean sd q0.025 q0.500 q0.975
beta[1] 0.031 0.186 -0.234 -0.009 0.456
beta[2] 0.463 0.13 0.168 0.524 0.646
beta[3] 0.41 0.209 0.103 0.461 0.82
beta[4] -0.185 0.075 -0.281 -0.117 -0.116
beta[5] 0.235 0.305 -0.42 0.247 0.74
-- Outcome fits --
[control]
MixGPD summary | backend: Stick-Breaking Process | kernel: Laplace Distribution | GPD tail: FALSE | epsilon: 0.025
n = 232 | components = 3
Summary
Initial components: 3 | Components after truncation: 3
Summary table
parameter mean sd q0.025 q0.500 q0.975
weights[1] 0.377 0.018 0.346 0.37 0.405
weights[2] 0.332 0.023 0.298 0.328 0.358
weights[3] 0.291 0.017 0.272 0.29 0.326
alpha 1.233 0.723 0.491 0.986 3.16
scale[1] 0.805 0.101 0.648 0.804 1.061
scale[2] 0.783 0.119 0.596 0.777 1.082
scale[3] 0.748 0.121 0.521 0.727 1.022
beta_location[1,1] 0 0 0 0 0
beta_location[1,2] 0 0 0 0 0
beta_location[1,3] 0 0 0 0 0
beta_location[1,4] 0 0 0 0 0
beta_location[2,1] 0 0 0 0 0
beta_location[2,2] 0 0 0 0 0
beta_location[2,3] 0 0 0 0 0
beta_location[2,4] 0 0 0 0 0
beta_location[3,1] 0 0 0 0 0
beta_location[3,2] 0 0 0 0 0
beta_location[3,3] 0 0 0 0 0
beta_location[3,4] 0 0 0 0 0
[treated]
MixGPD summary | backend: Stick-Breaking Process | kernel: Laplace Distribution | GPD tail: FALSE | epsilon: 0.025
n = 268 | components = 3
Summary
Initial components: 3 | Components after truncation: 3
Summary table
parameter mean sd q0.025 q0.500 q0.975
weights[1] 0.698 0.017 0.679 0.713 0.713
weights[2] 0.161 0.013 0.144 0.162 0.186
weights[3] 0.141 0.021 0.101 0.143 0.16
alpha 1.069 0.548 0.25 0.888 2.367
scale[1] 0.685 0.073 0.568 0.677 0.838
scale[2] 0.675 0.184 0.469 0.632 1.239
scale[3] 0.674 0.154 0.433 0.654 1.03
beta_location[1,1] 0 0 0 0 0
beta_location[1,2] 0 0 0 0 0
beta_location[1,3] 0 0 0 0 0
beta_location[1,4] 0 0 0 0 0
beta_location[2,1] 0 0 0 0 0
beta_location[2,2] 0 0 0 0 0
beta_location[2,3] 0 0 0 0 0
beta_location[2,4] 0 0 0 0 0
beta_location[3,1] 0 0 0 0 0
beta_location[3,2] 0 0 0 0 0
beta_location[3,3] 0 0 0 0 0
beta_location[3,4] 0 0 0 0 0
Code
Posterior mean parameters (causal)
[ps]
Posterior mean parameters
$beta
[1] 0.03087 0.46270 0.40980 -0.18530 0.23550
[treated]
Posterior mean parameters
$alpha
[1] 1.069
$w
[1] 0.6980 0.1607 0.1413
$beta_location
PropScore x1 x2 x3 x4
comp1 0 0 0 0 0
comp2 0 0 0 0 0
comp3 0 0 0 0 0
$scale
[1] 0.6849 0.6750 0.6736
[control]
Posterior mean parameters
$alpha
[1] 1.233
$w
[1] 0.3765 0.3322 0.2913
$beta_location
PropScore x1 x2 x3 x4
comp1 0 0 0 0 0
comp2 0 0 0 0 0
comp3 0 0 0 0 0
$scale
[1] 0.8050 0.7825 0.7482
Code
plot (fit_sb_bulk, family = plot_family_bulk_ex12)
=== treated ===
=== traceplot ===
=== control ===
=== traceplot ===
Code
<- predict (fit_sb_bulk, newdata = x_eval, type = "mean" ,interval = "credible" , nsim_mean = nsim_mean_ex12, workers = predict_workers_ex12)plot (pred_mean_bulk)
Code
<- predict (fit_sb_bulk, newdata = x_eval, type = "quantile" ,p = 0.5 , interval = "credible" , workers = predict_workers_ex12)plot (pred_q_bulk)
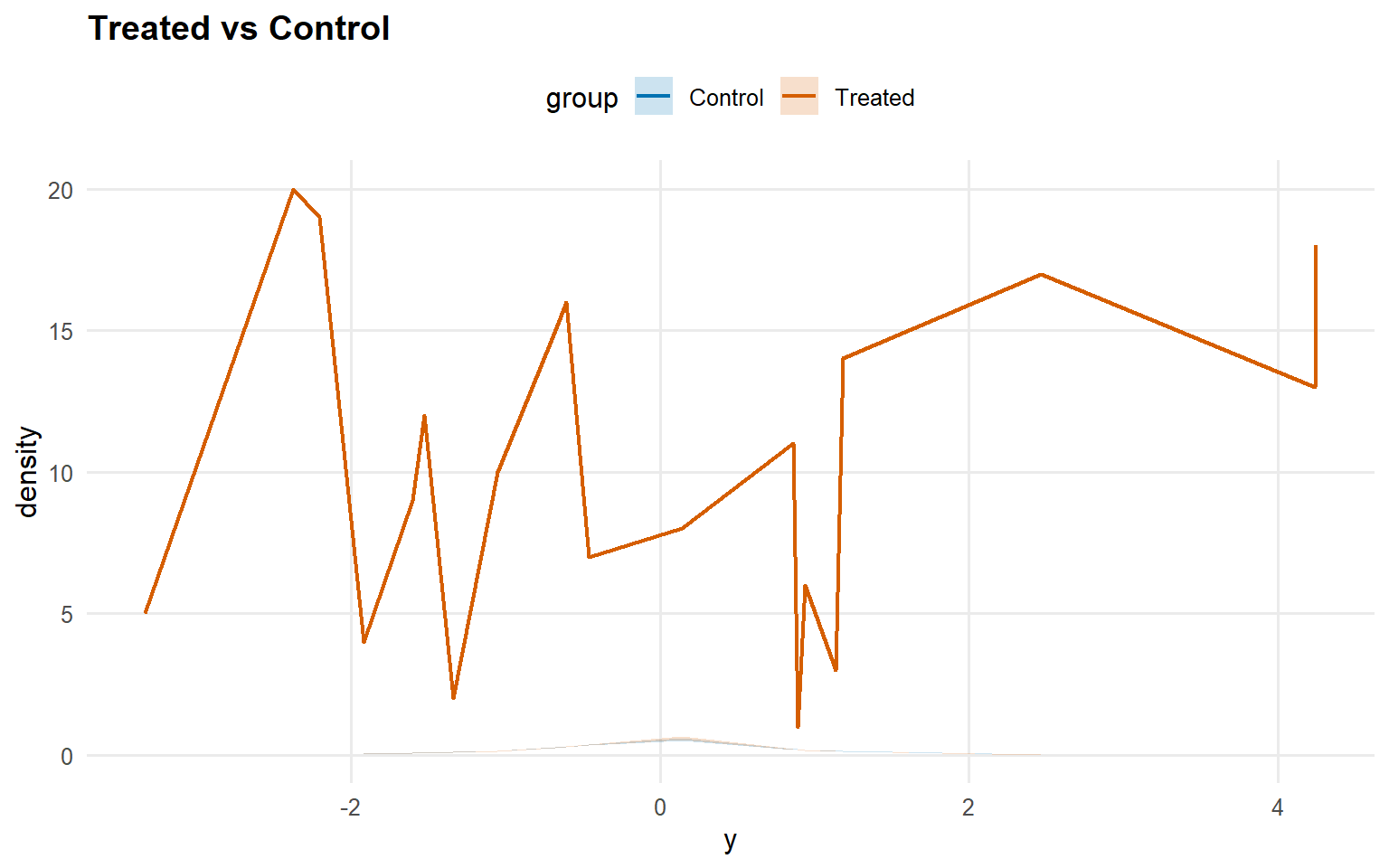
Code
<- predict (fit_sb_bulk, newdata = x_eval, y = y_eval,type = "density" , interval = "credible" , workers = predict_workers_ex12)plot (pred_d_bulk)
Code
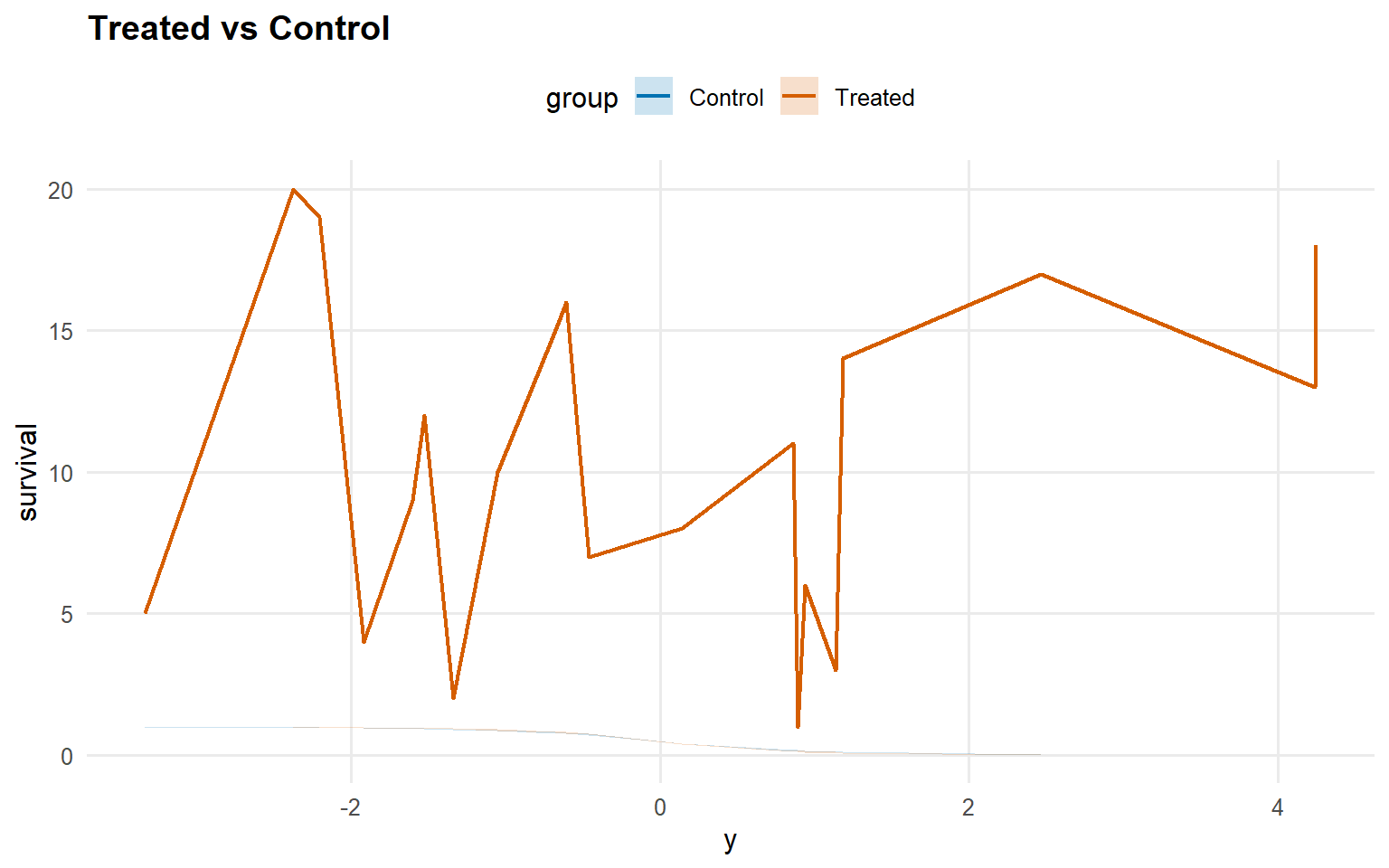
<- predict (fit_sb_bulk, newdata = x_eval, y = y_eval,type = "survival" , interval = "credible" , workers = predict_workers_ex12)plot (pred_surv_bulk)
Code
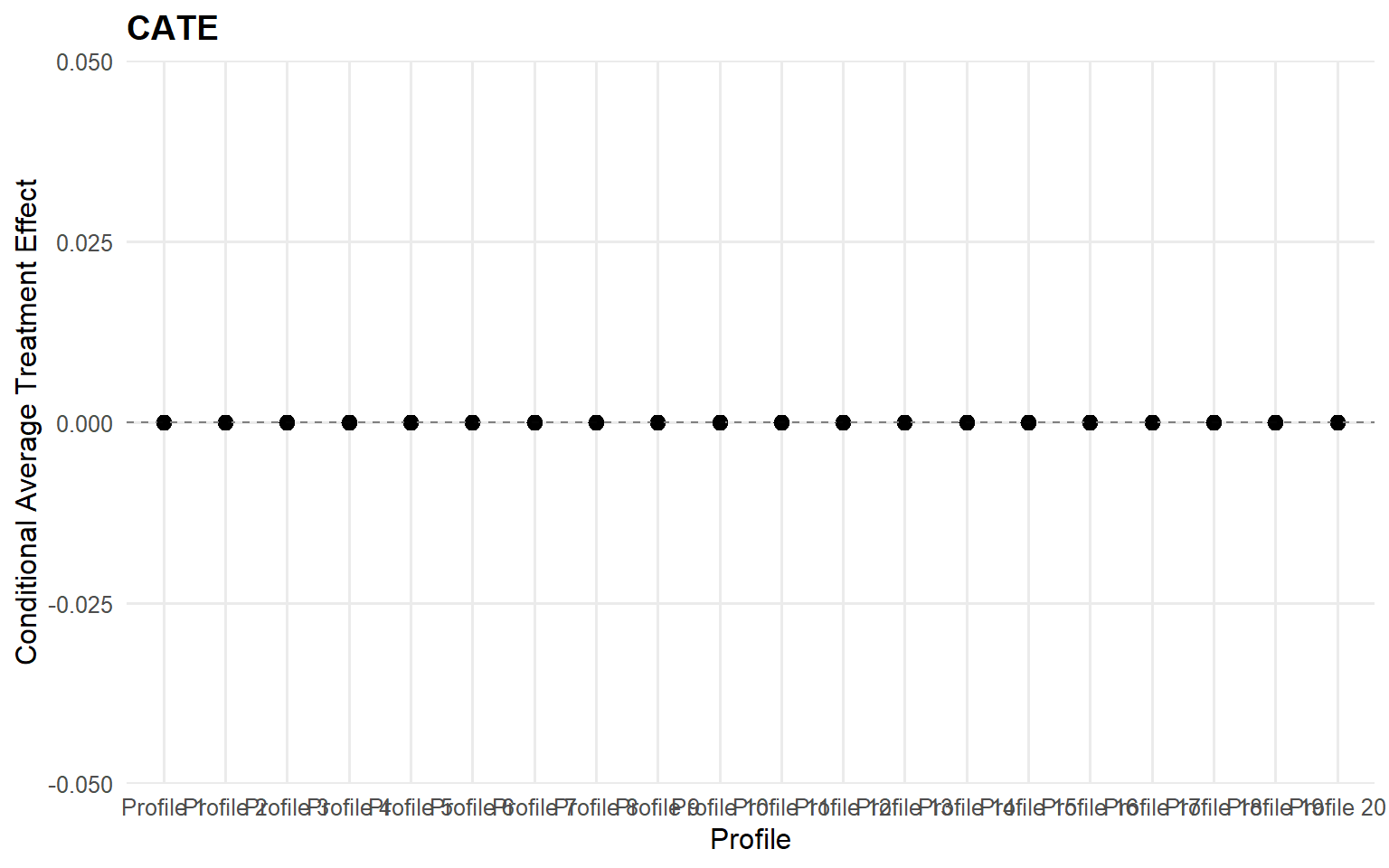
<- cate (fit_sb_bulk, newdata = x_eval,interval = "credible" , nsim_mean = nsim_mean_ex12)head (format_cate_table (cate_bulk))
id index estimate lower upper
1 1 1 0 0 0
2 2 1 0 0 0
3 3 1 0 0 0
4 4 1 0 0 0
5 5 1 0 0 0
6 6 1 0 0 0
Code
Code
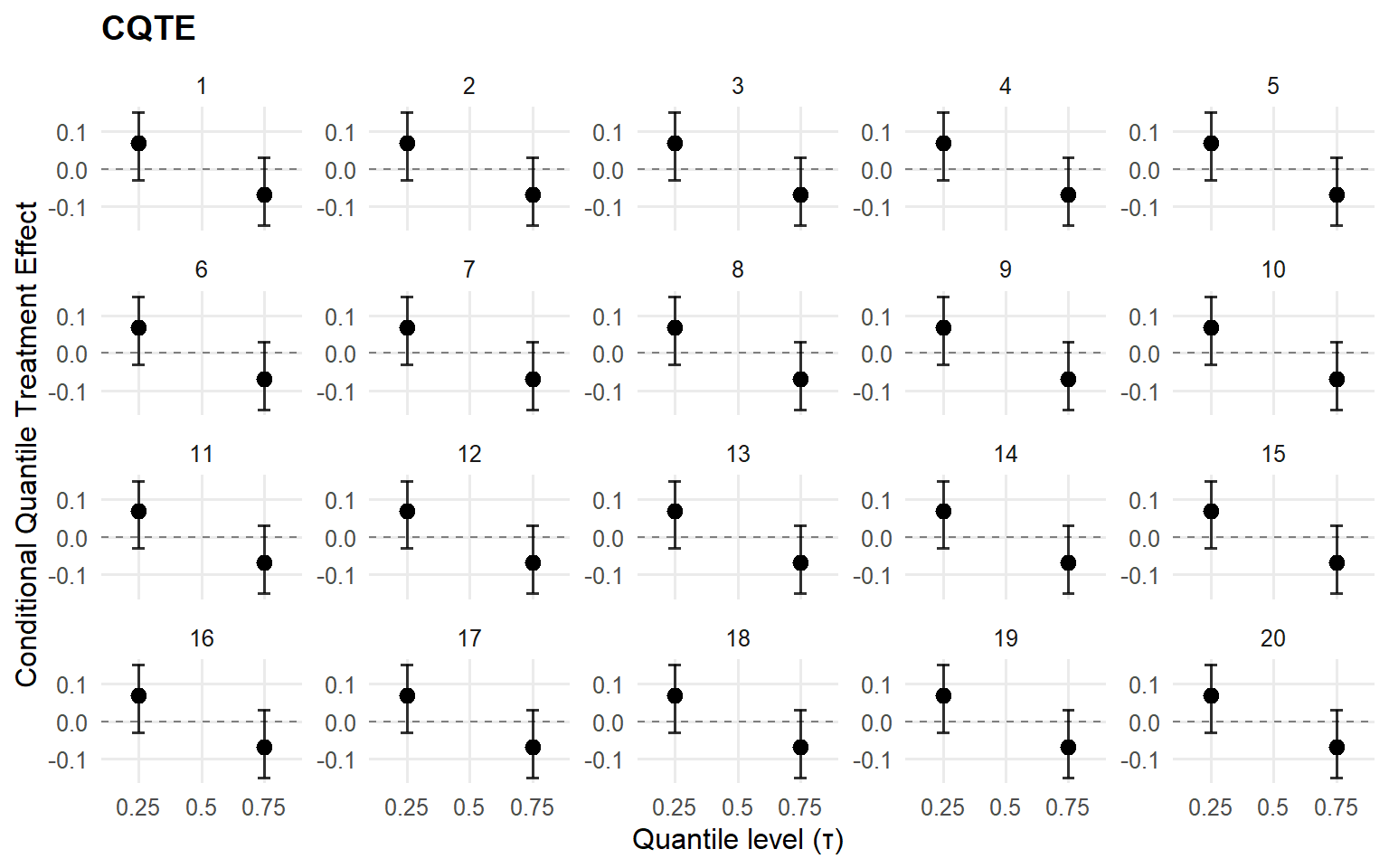
<- cqte (fit_sb_bulk, probs = c (0.25 , 0.5 , 0.75 ),newdata = x_eval, interval = "credible" )head (format_cqte_table (cqte_bulk))
id index estimate lower upper
1 1 1 0.06878438 -0.0300103 0.1503468
2 1 2 NaN NA NA
3 1 3 -0.06878438 -0.1503468 0.0300103
4 2 1 0.06878438 -0.0300103 0.1503468
5 2 2 NaN NA NA
6 2 3 -0.06878438 -0.1503468 0.0300103
Code
Model B: SB with GPD Tail (Laplace)
Code
<- list (gpd = list (threshold = list (mode = "dist" ,dist = "lognormal" ,args = list (meanlog = log (max (u_threshold, .Machine$ double.eps)), sdlog = 0.25 )<- bundle (y = y,treat = A,X = X,kernel = "laplace" ,backend = "sb" ,PS = "logit" ,GPD = TRUE ,components = components_ex12,param_specs = param_specs_gpd,mcmc_outcome = gpd_mcmc_ex12,mcmc_ps = mcmc_ps_gpd_ex12
CausalMixGPD causal bundle
PS model: Bayesian logit (A | X)
Field Treated Control
Backend Stick-Breaking Process Stick-Breaking Process
Kernel laplace laplace
Components 3 3
GPD tail TRUE TRUE
Epsilon 0.025 0.025
Outcome PS included: TRUE
n (control) = 232 | n (treated) = 268
Code
CausalMixGPD causal bundle summary
CausalMixGPD causal bundle
PS model: Bayesian logit (A | X)
Field Treated Control
Backend Stick-Breaking Process Stick-Breaking Process
Kernel laplace laplace
Components 3 3
GPD tail TRUE TRUE
Epsilon 0.025 0.025
Outcome PS included: TRUE
n (control) = 232 | n (treated) = 268
Code
<- load_or_fit (quiet_mcmc (dpmgpd.causal (bundle_sb_gpd, mcmc = causal_run_mcmc_gpd))summary (fit_sb_gpd)
-- PS fit --
CausalMixGPD PS fit summary
model: logit
n = 500 | predictors = 5
Monitors: beta
Summary table
parameter mean sd q0.025 q0.500 q0.975
beta[1] 0.021 0.082 -0.056 -0.005 0.144
beta[2] 0.458 0.082 0.324 0.451 0.607
beta[3] 0.27 0.197 -0.042 0.216 0.544
beta[4] -0.149 0.065 -0.303 -0.107 -0.07
beta[5] 0.277 0.149 -0.228 0.348 0.48
-- Outcome fits --
[control]
MixGPD summary | backend: Stick-Breaking Process | kernel: Laplace Distribution | GPD tail: TRUE | epsilon: 0.025
n = 232 | components = 3
Summary
Initial components: 3 | Components after truncation: 3
Summary table
parameter mean sd q0.025 q0.500 q0.975
weights[1] 0.665 0.002 0.662 0.666 0.666
weights[2] 0.281 0.013 0.264 0.288 0.302
weights[3] 0.054 0.015 0.035 0.046 0.074
alpha 1.094 0.551 0.475 1.027 2.832
beta_tail_scale[1] 0.545 0.26 0.116 0.56 0.996
beta_tail_scale[2] 0.078 0.321 -0.472 0.12 0.648
beta_tail_scale[3] -0.019 0.262 -0.44 -0.009 0.507
beta_tail_scale[4] -0.555 0.397 -1.214 -0.589 0.15
tail_shape 0.182 0.188 -0.286 0.176 0.414
threshold 2.427 0.088 2.151 2.46 2.46
scale[1] 1.43 0.15 1.186 1.443 1.719
scale[2] 1.502 0.247 1.123 1.466 1.958
scale[3] 1.889 0.989 0.644 1.646 4.361
beta_location[1,1] 0 0 0 0 0
beta_location[1,2] 0 0 0 0 0
beta_location[1,3] 0 0 0 0 0
beta_location[1,4] 0 0 0 0 0
beta_location[2,1] 0 0 0 0 0
beta_location[2,2] 0 0 0 0 0
beta_location[2,3] 0 0 0 0 0
beta_location[2,4] 0 0 0 0 0
beta_location[3,1] 0 0 0 0 0
beta_location[3,2] 0 0 0 0 0
beta_location[3,3] 0 0 0 0 0
beta_location[3,4] 0 0 0 0 0
[treated]
MixGPD summary | backend: Stick-Breaking Process | kernel: Laplace Distribution | GPD tail: TRUE | epsilon: 0.025
n = 268 | components = 3
Summary
Initial components: 3 | Components after truncation: 3
Summary table
parameter mean sd q0.025 q0.500 q0.975
weights[1] 0.452 0.038 0.437 0.437 0.551
weights[2] 0.321 0.026 0.285 0.32 0.361
weights[3] 0.227 0.035 0.161 0.243 0.278
alpha 1.447 0.968 0.206 1.282 3.431
beta_tail_scale[1] 0.154 0.256 -0.278 0.118 0.734
beta_tail_scale[2] 0.063 0.369 -0.463 0.011 0.75
beta_tail_scale[3] 0.149 0.162 -0.143 0.152 0.478
beta_tail_scale[4] 0.038 0.536 -0.842 -0.018 1.333
tail_shape -0.144 0.269 -0.583 -0.209 0.226
threshold 2.306 1.025 1.069 2.001 4.037
scale[1] 1.68 0.253 1.339 1.645 2.287
scale[2] 1.73 0.29 1.242 1.708 2.319
scale[3] 1.585 0.511 1.062 1.398 3.161
beta_location[1,1] 0 0 0 0 0
beta_location[1,2] 0 0 0 0 0
beta_location[1,3] 0 0 0 0 0
beta_location[1,4] 0 0 0 0 0
beta_location[2,1] 0 0 0 0 0
beta_location[2,2] 0 0 0 0 0
beta_location[2,3] 0 0 0 0 0
beta_location[2,4] 0 0 0 0 0
beta_location[3,1] 0 0 0 0 0
beta_location[3,2] 0 0 0 0 0
beta_location[3,3] 0 0 0 0 0
beta_location[3,4] 0 0 0 0 0
Code
Posterior mean parameters (causal)
[ps]
Posterior mean parameters
$beta
[1] 0.02086 0.45850 0.27050 -0.14870 0.27740
[treated]
Posterior mean parameters
$alpha
[1] 1.447
$w
[1] 0.4521 0.3205 0.2274
$beta_location
PropScore x1 x2 x3 x4
comp1 0 0 0 0 0
comp2 0 0 0 0 0
comp3 0 0 0 0 0
$scale
[1] 1.680 1.730 1.585
$threshold
[1] 2.306
$beta_tail_scale
x1 x2 x3 x4
overall 0.1536 0.0633 0.149 0.03797
$tail_shape
[1] -0.1438
[control]
Posterior mean parameters
$alpha
[1] 1.094
$w
[1] 0.66460 0.28140 0.05398
$beta_location
PropScore x1 x2 x3 x4
comp1 0 0 0 0 0
comp2 0 0 0 0 0
comp3 0 0 0 0 0
$scale
[1] 1.430 1.502 1.889
$threshold
[1] 2.427
$beta_tail_scale
x1 x2 x3 x4
overall 0.5454 0.0782 -0.01914 -0.5554
$tail_shape
[1] 0.1823
Code
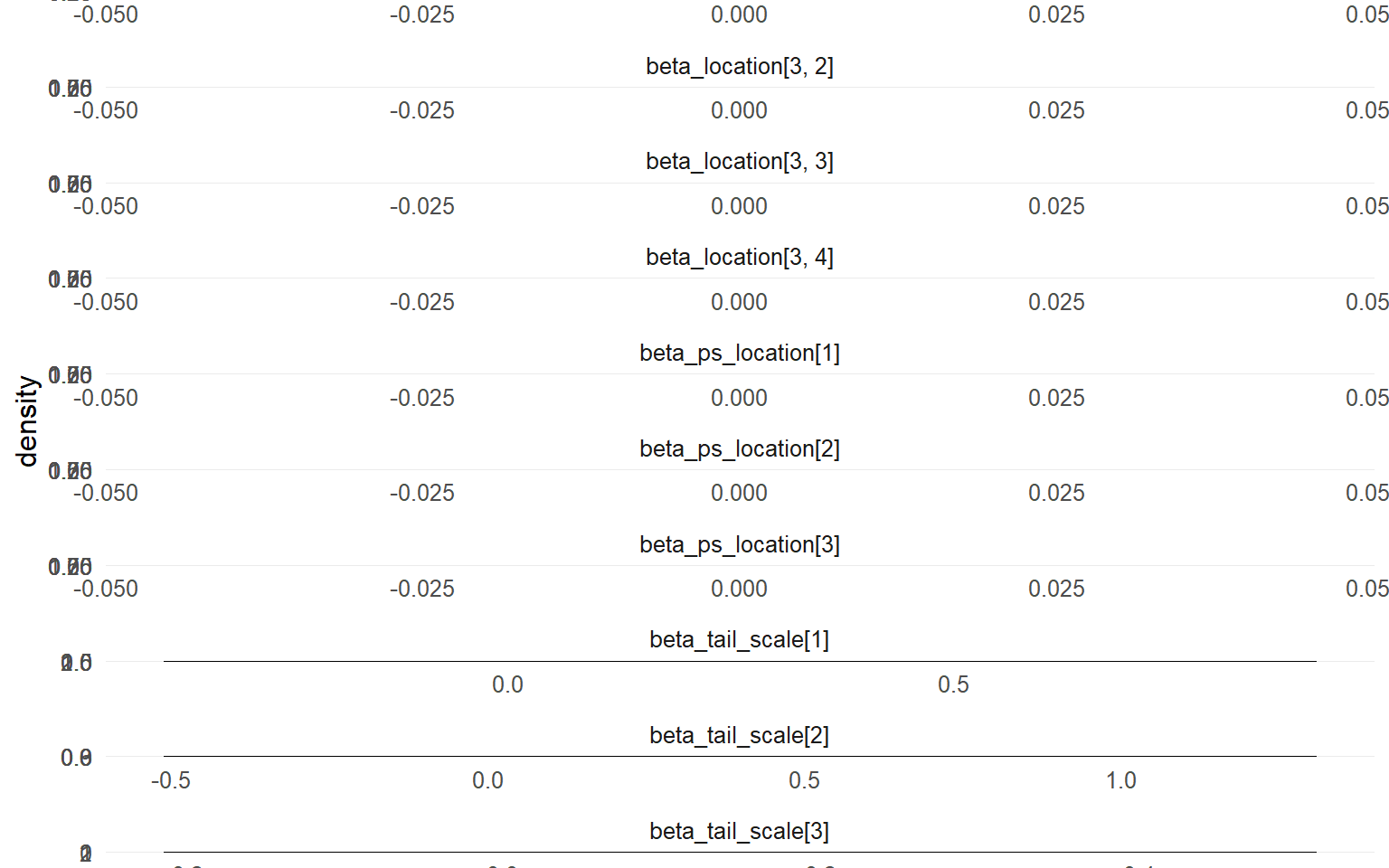
plot (fit_sb_gpd, family = plot_family_gpd_ex12)
=== treated ===
=== density ===
=== control ===
=== density ===
Code
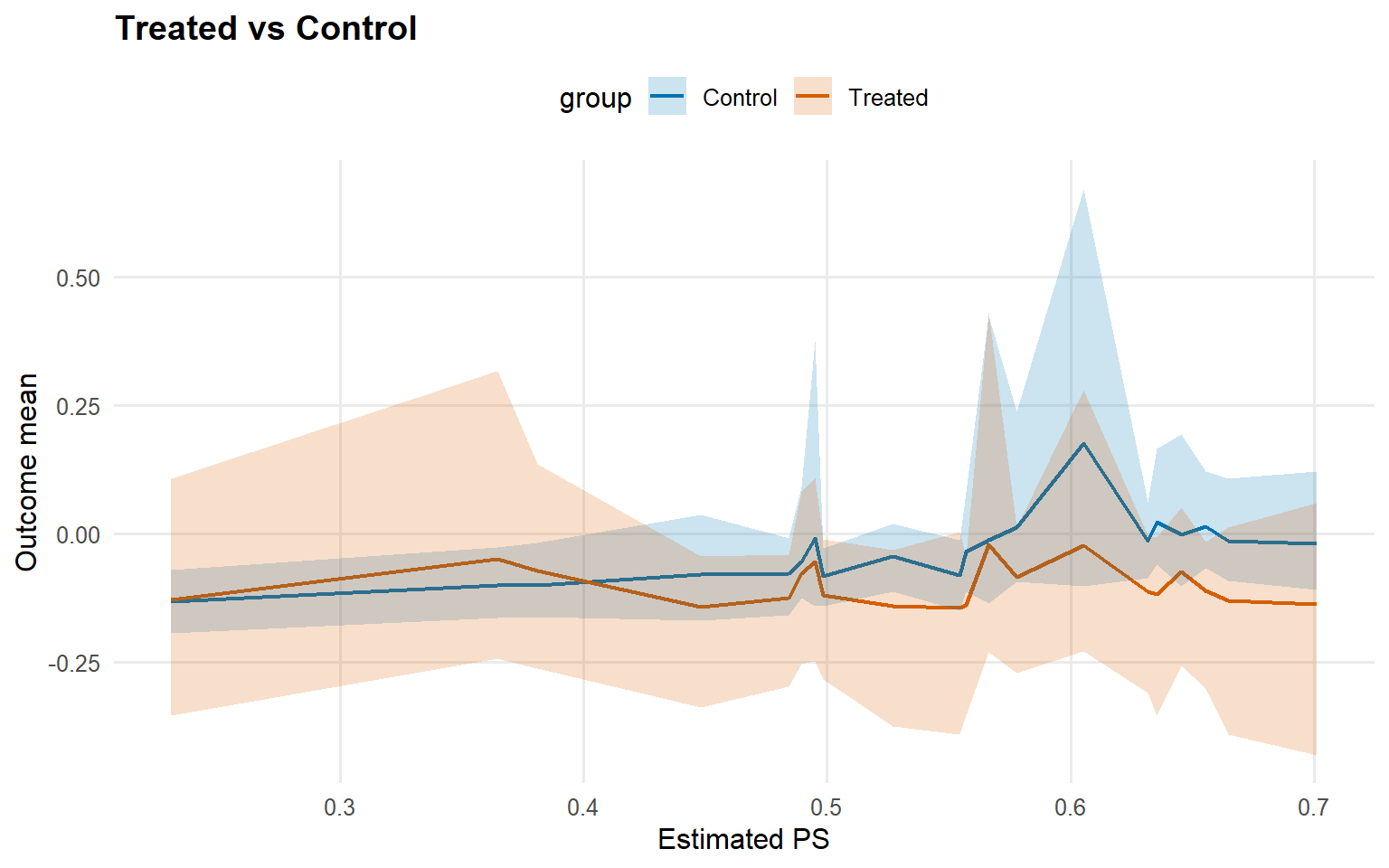
<- predict (fit_sb_gpd, newdata = x_eval, type = "mean" ,interval = "credible" , nsim_mean = nsim_mean_ex12, workers = predict_workers_ex12)plot (pred_mean_gpd)
Code
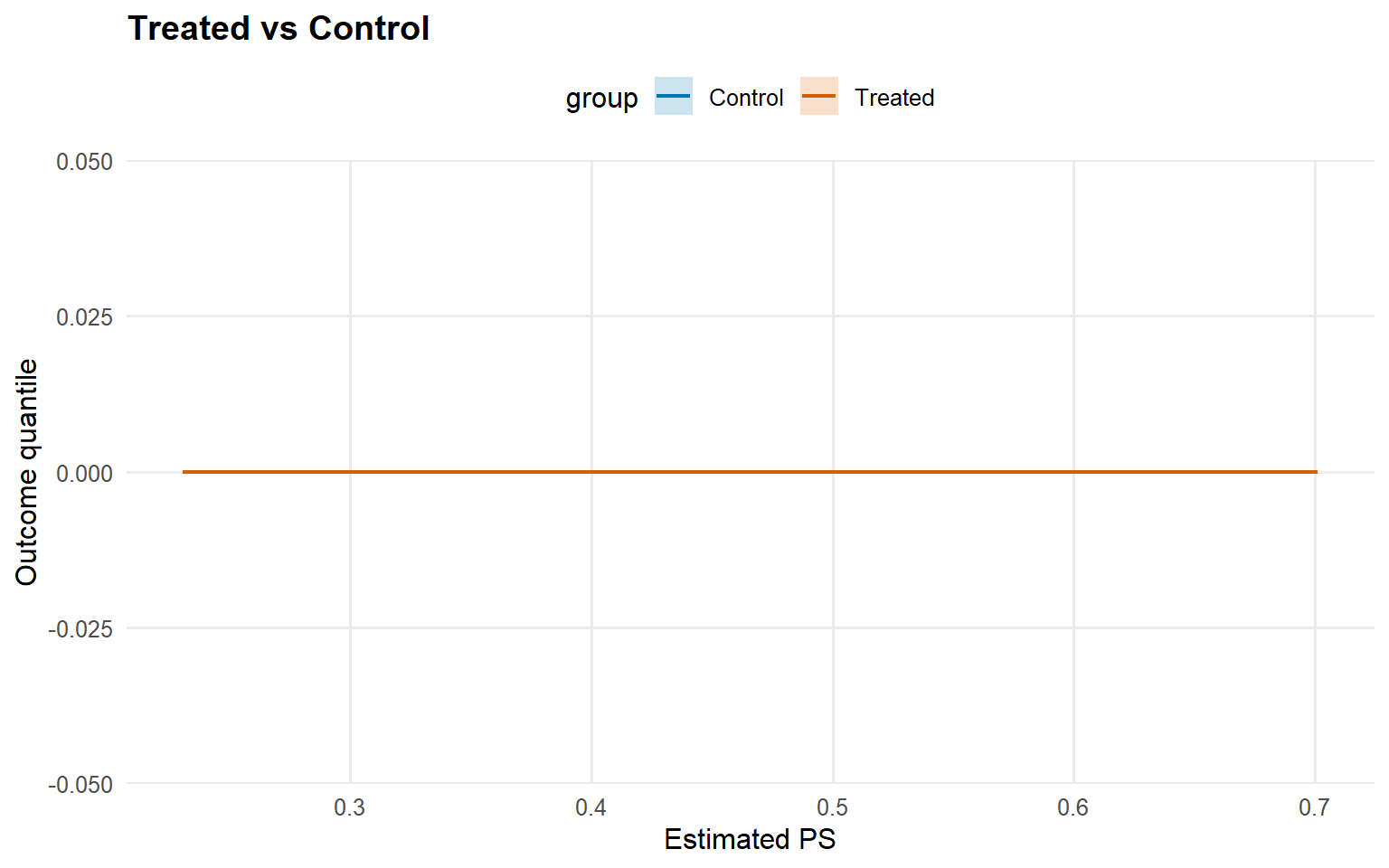
<- predict (fit_sb_gpd, newdata = x_eval, type = "quantile" ,p = 0.5 , interval = "credible" , workers = predict_workers_ex12)plot (pred_q_gpd)
Code
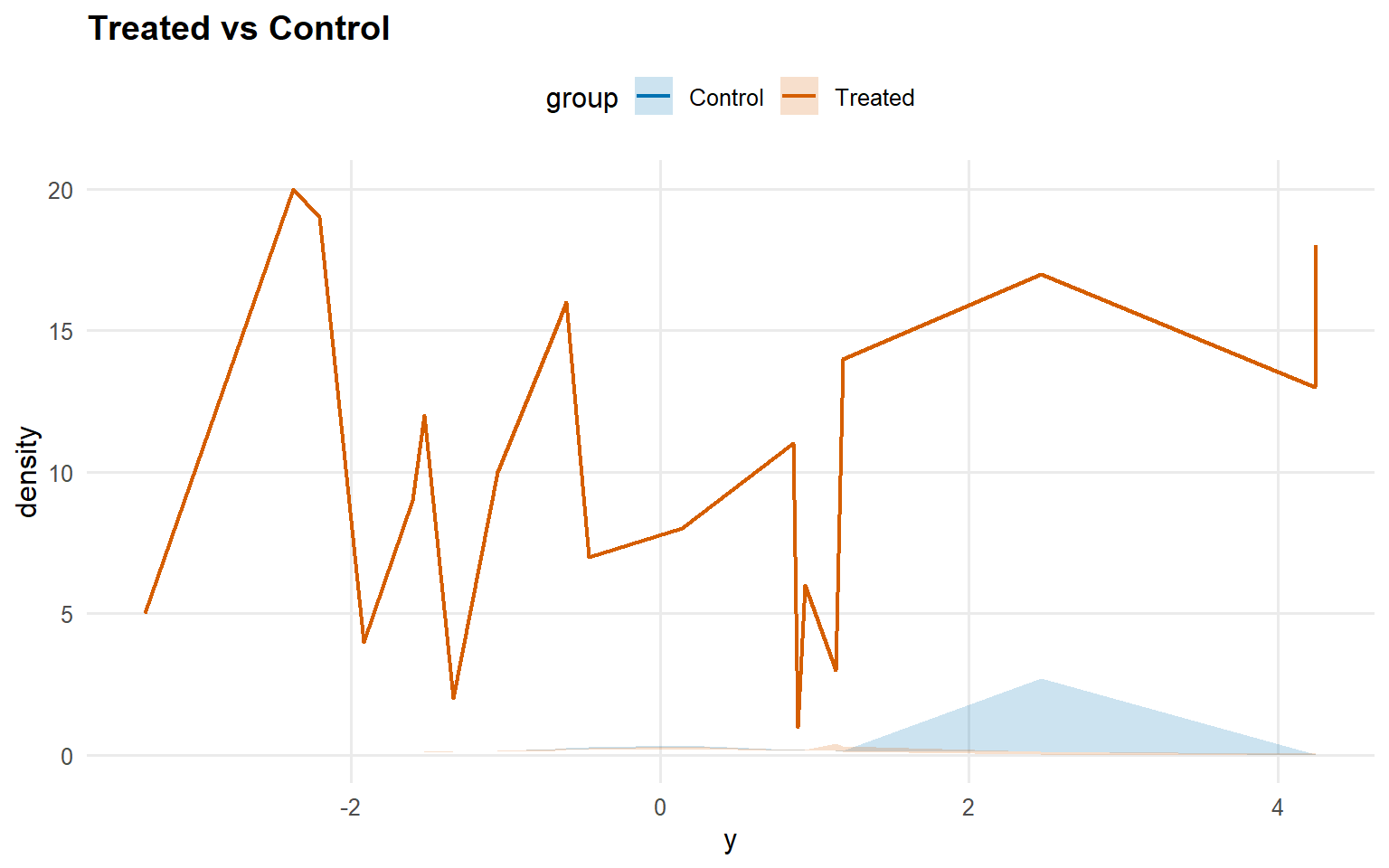
<- predict (fit_sb_gpd, newdata = x_eval, y = y_eval,type = "density" , interval = "credible" , workers = predict_workers_ex12)plot (pred_d_gpd)
Code
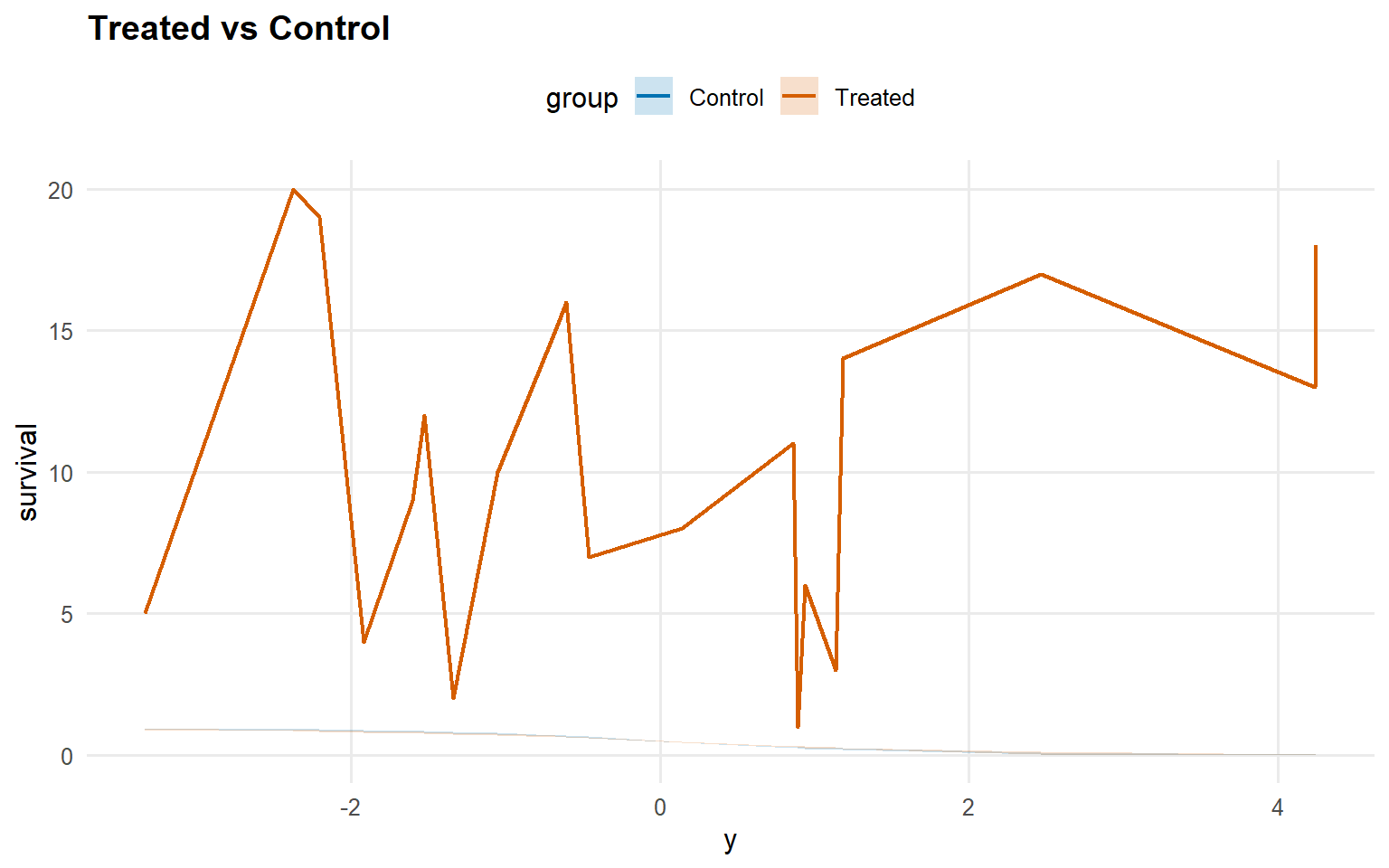
<- predict (fit_sb_gpd, newdata = x_eval, y = y_eval,type = "survival" , interval = "credible" , workers = predict_workers_ex12)plot (pred_surv_gpd)
Code
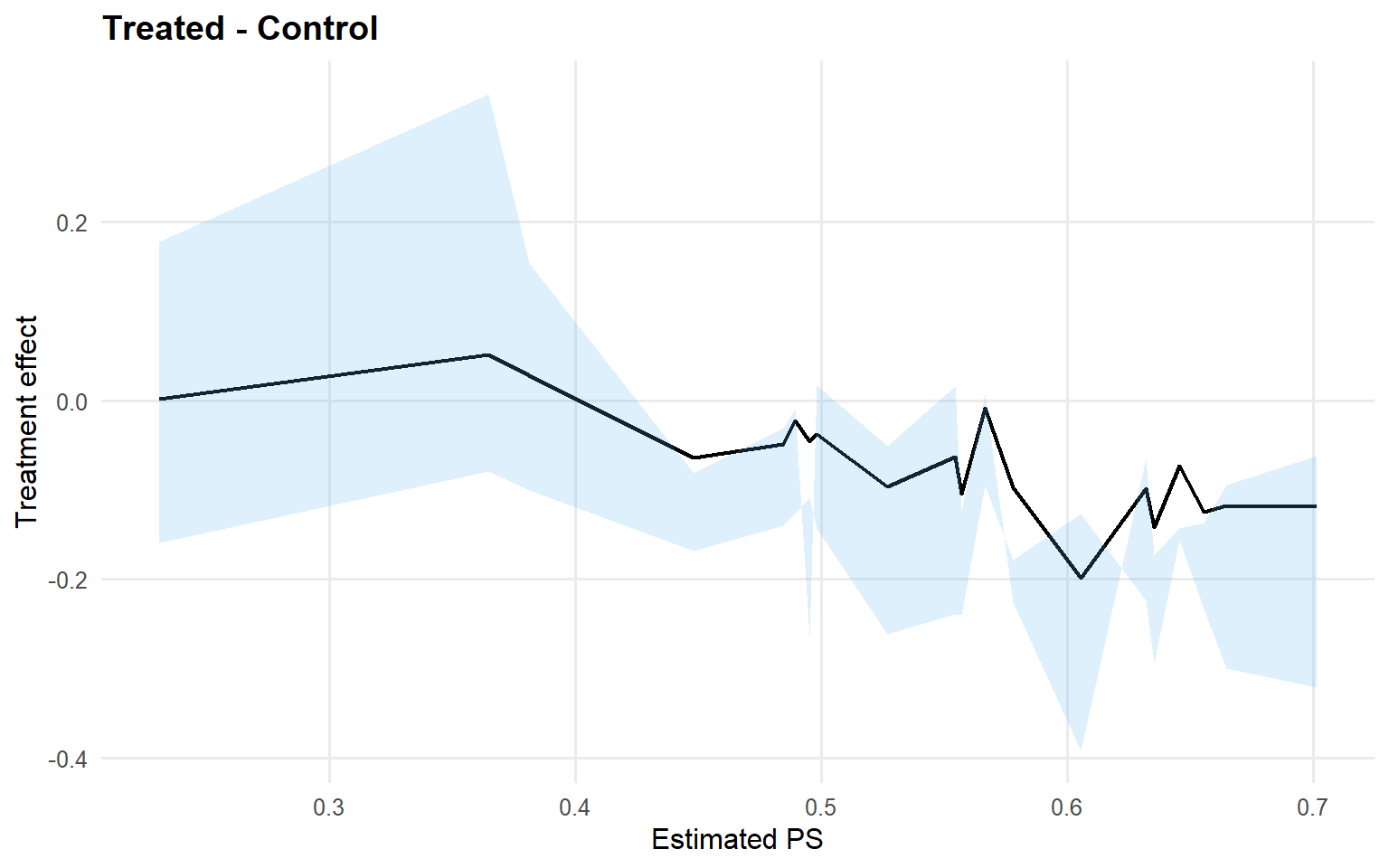
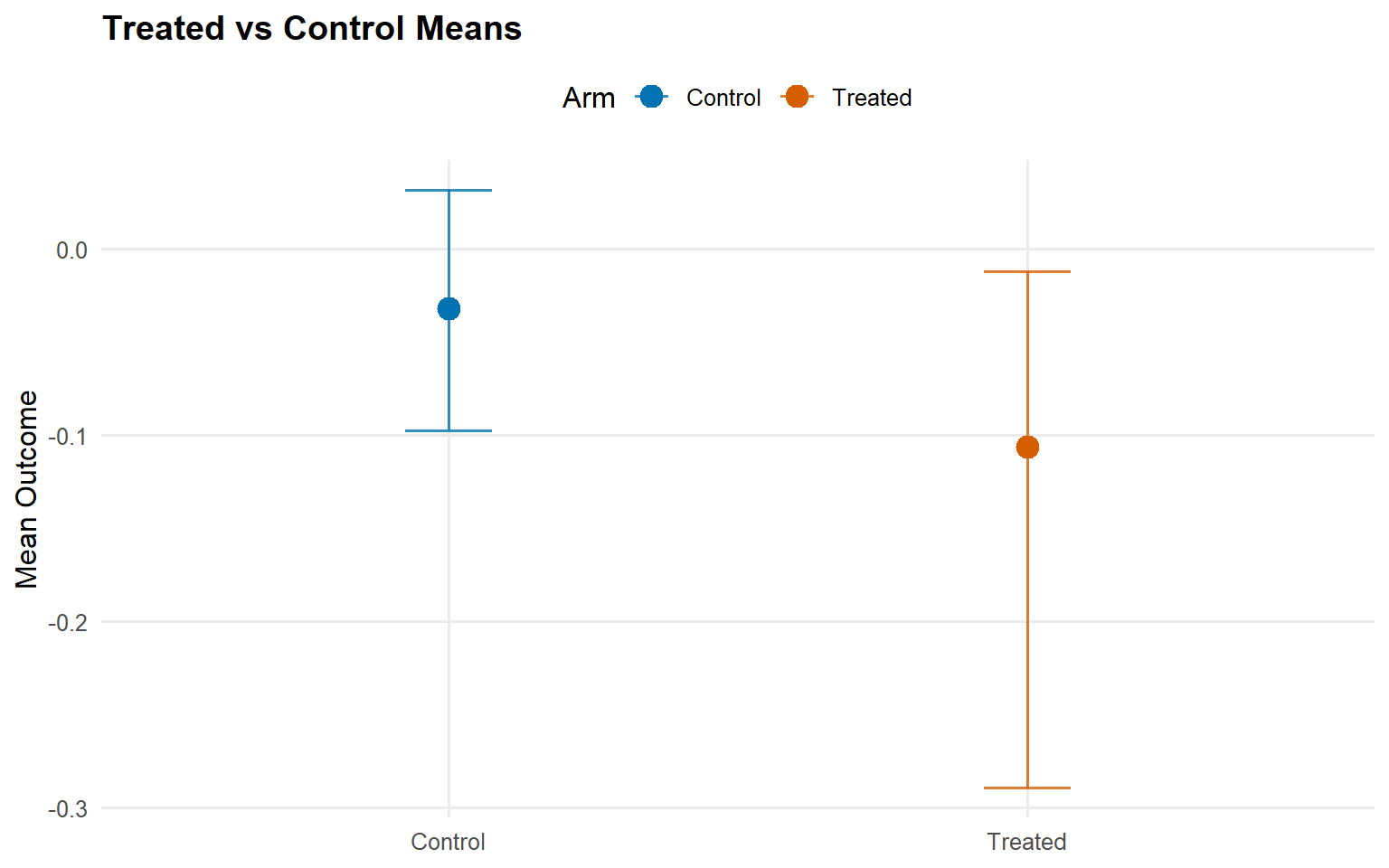
<- ate (fit_sb_gpd, interval = "credible" , nsim_mean = nsim_mean_ex12)plot (ate_gpd)
Code
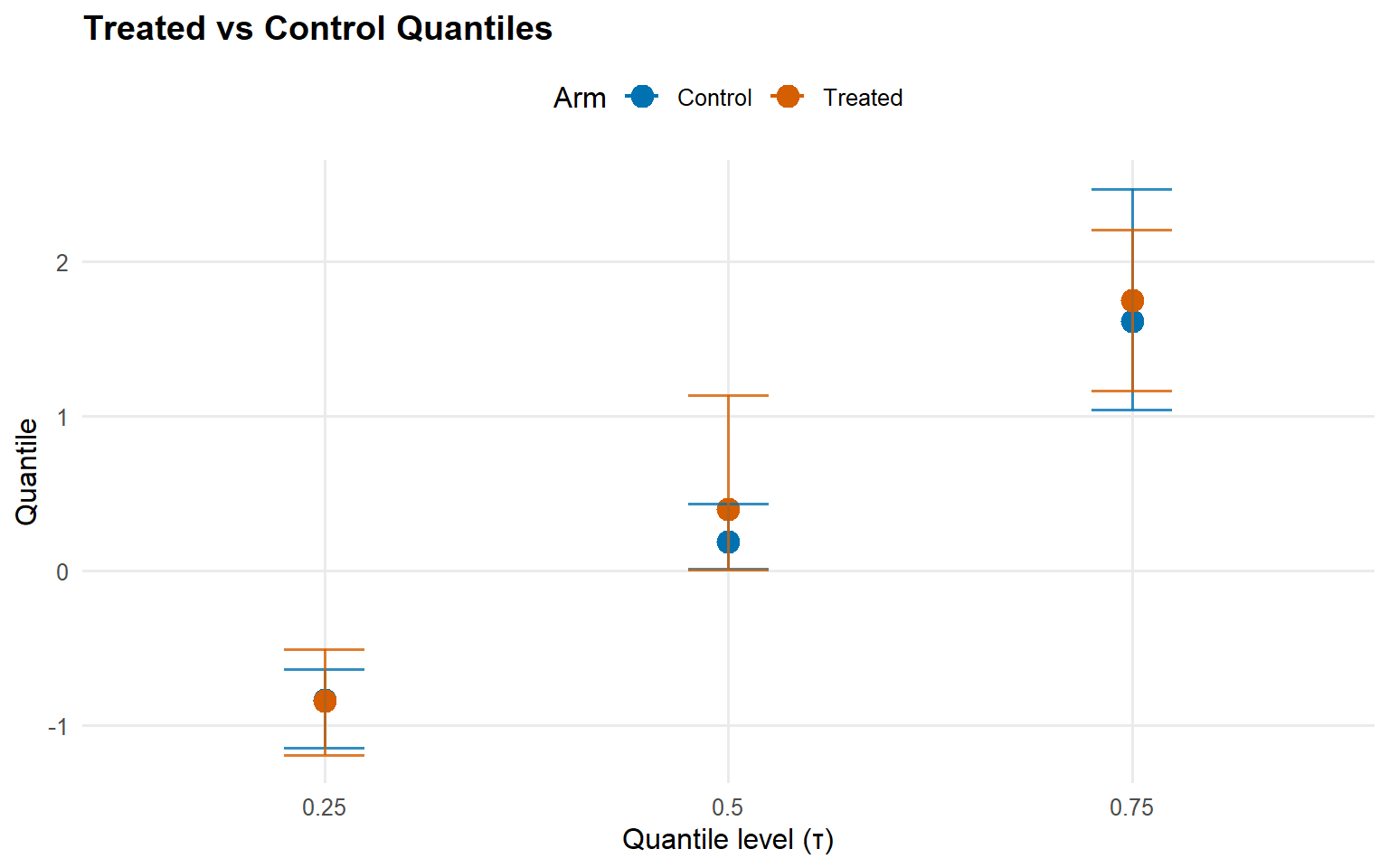
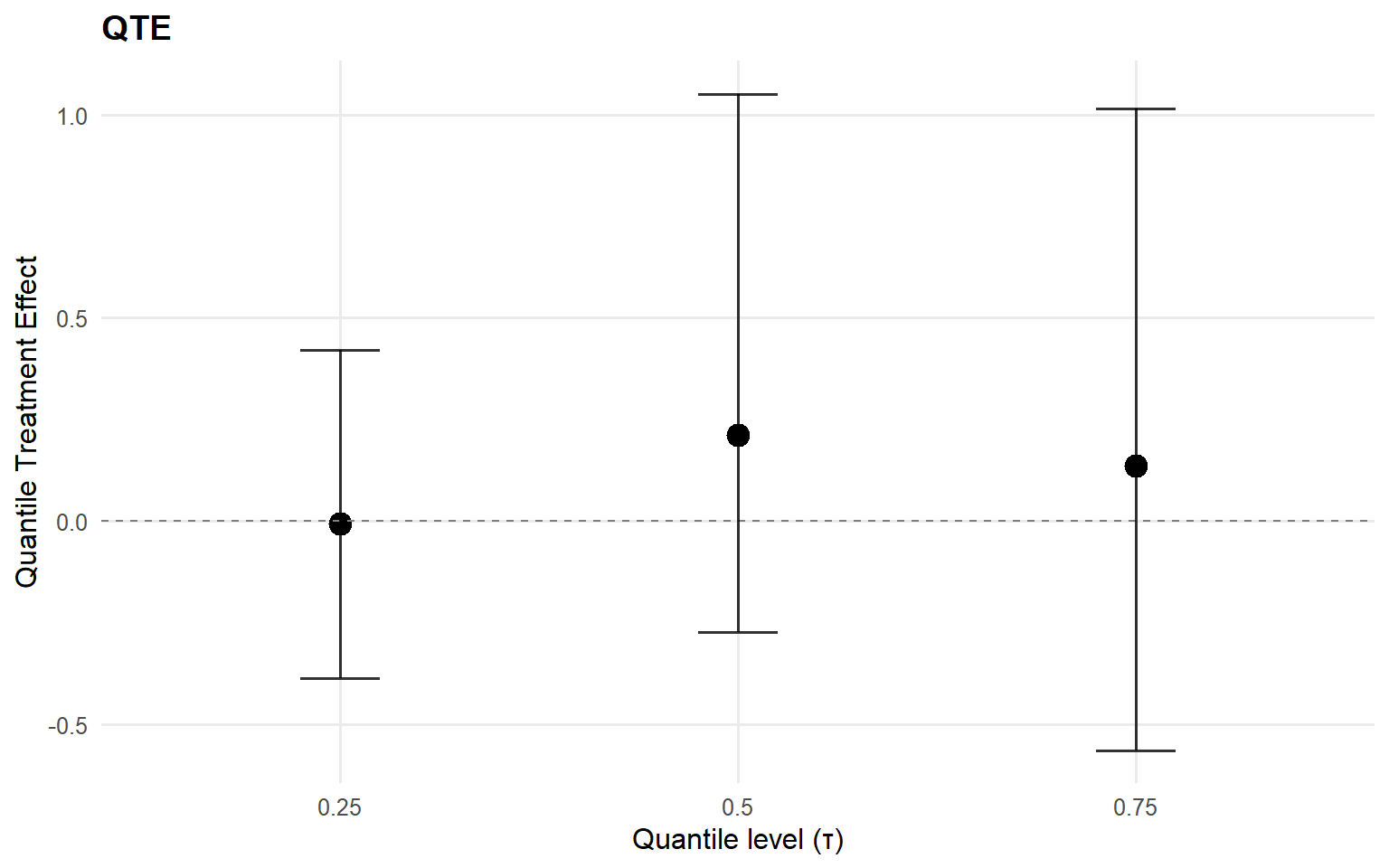
<- qte (fit_sb_gpd, probs = c (0.25 , 0.5 , 0.75 ), interval = "credible" )plot (qte_gpd)
Prereqs
Required packages and data for this page are listed in the setup chunks above.
Outputs
This page renders model fits, diagnostics, and summary artifacts generated by package APIs.
Interpretation
Canonical concept page: 03 Causal Inference Objects
Treat this page as an application/example view and use the canonical page for core definitions.
Next
Continue to the linked canonical concept page, then return for implementation-specific details.