Website workflow note. This page reflects the current exported API and recommended wrapper-first usage. Last updated: 2026-02-19 .
For the full package narrative, see the main package vignettes (basic, unconditional, conditional, and causal).
Causal Inference: Mixed Backends (Bulk-only) - Cauchy Kernel
This vignette fits two mixed-backend causal models with a shared Cauchy kernel:
Model A: Treated = SB, Control = CRP
Model B: Treated = CRP, Control = SB
What you’ll learn
How to specify arm-specific backends (treated vs control) within one causal bundle.
How to interpret causal predictions/effects when the two potential-outcome models have different partition structures.
How to keep the comparison focused on backend differences by using a shared kernel and GPD = FALSE.
When to use this template
You suspect treated and control outcomes differ in ways that make one backend preferable for one arm.
You want to study how much causal estimands change under backend asymmetry while keeping tail modeling off.
Next steps
Turn on tail augmentation (mixed-backend + GPD = TRUE) when extremes plausibly drive effect heterogeneity.
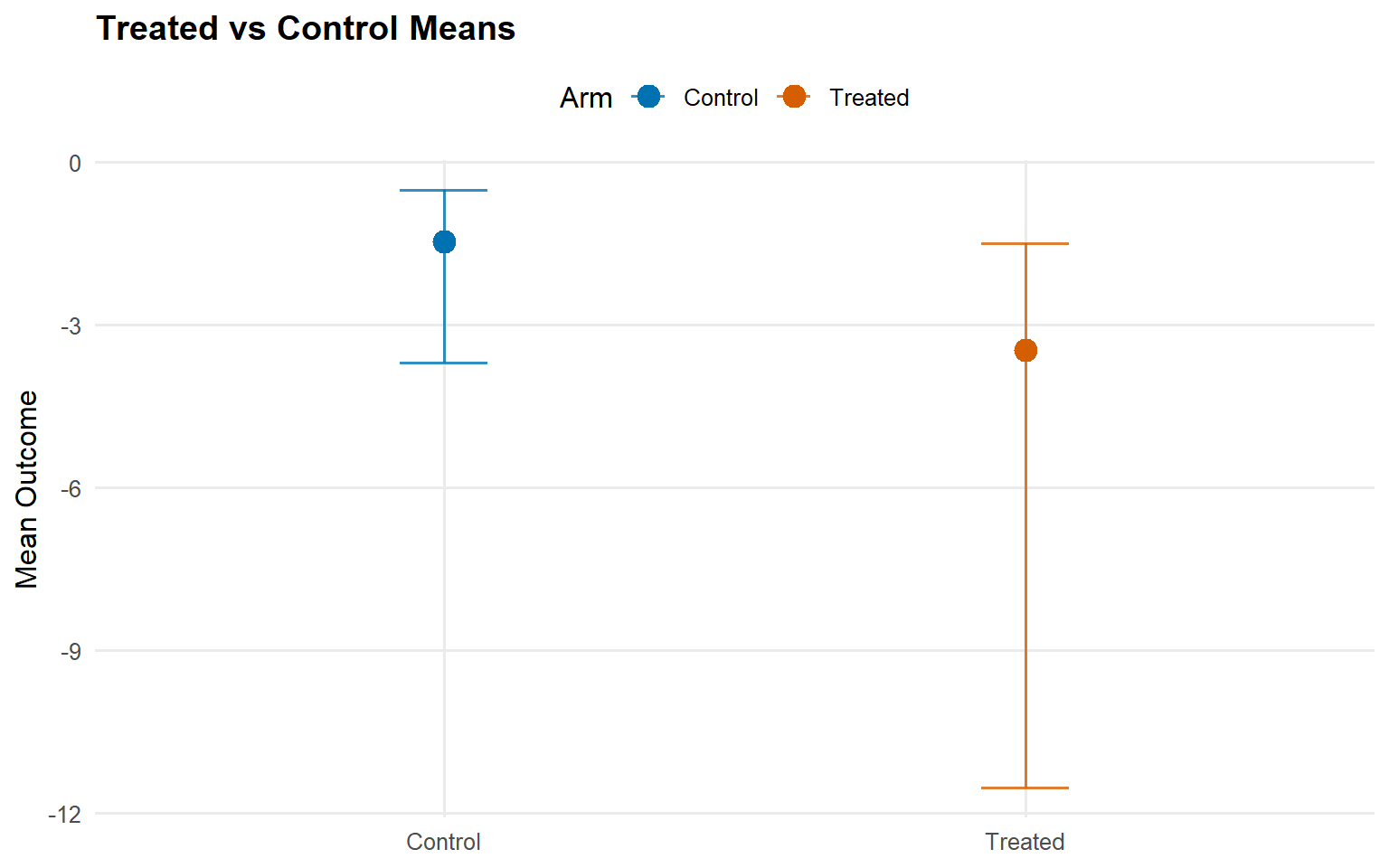
Because the Cauchy kernel does not have a finite mean, this page uses median and upper-quantile predictions, plus restricted-mean causal effects.
Data Setup
Code
data ("causal_alt_real500_p4_k2" )<- causal_alt_real500_p4_k2$ y<- causal_alt_real500_p4_k2$ A<- as.matrix (causal_alt_real500_p4_k2$ X)<- tibble (statistic = c ("N" , "Mean" , "SD" , "Min" , "Max" ),value = c (length (y), mean (y), sd (y), min (y), max (y))
# A tibble: 5 × 2
statistic value
<chr> <dbl>
1 N 500
2 Mean 0.274
3 SD 1.76
4 Min -8.09
5 Max 5.27
Code
<- X[seq_len (if (isTRUE (FAST)) 20 L else 40 L), , drop = FALSE ]<- y[seq_len (if (isTRUE (FAST)) 20 L else 40 L)]
Code
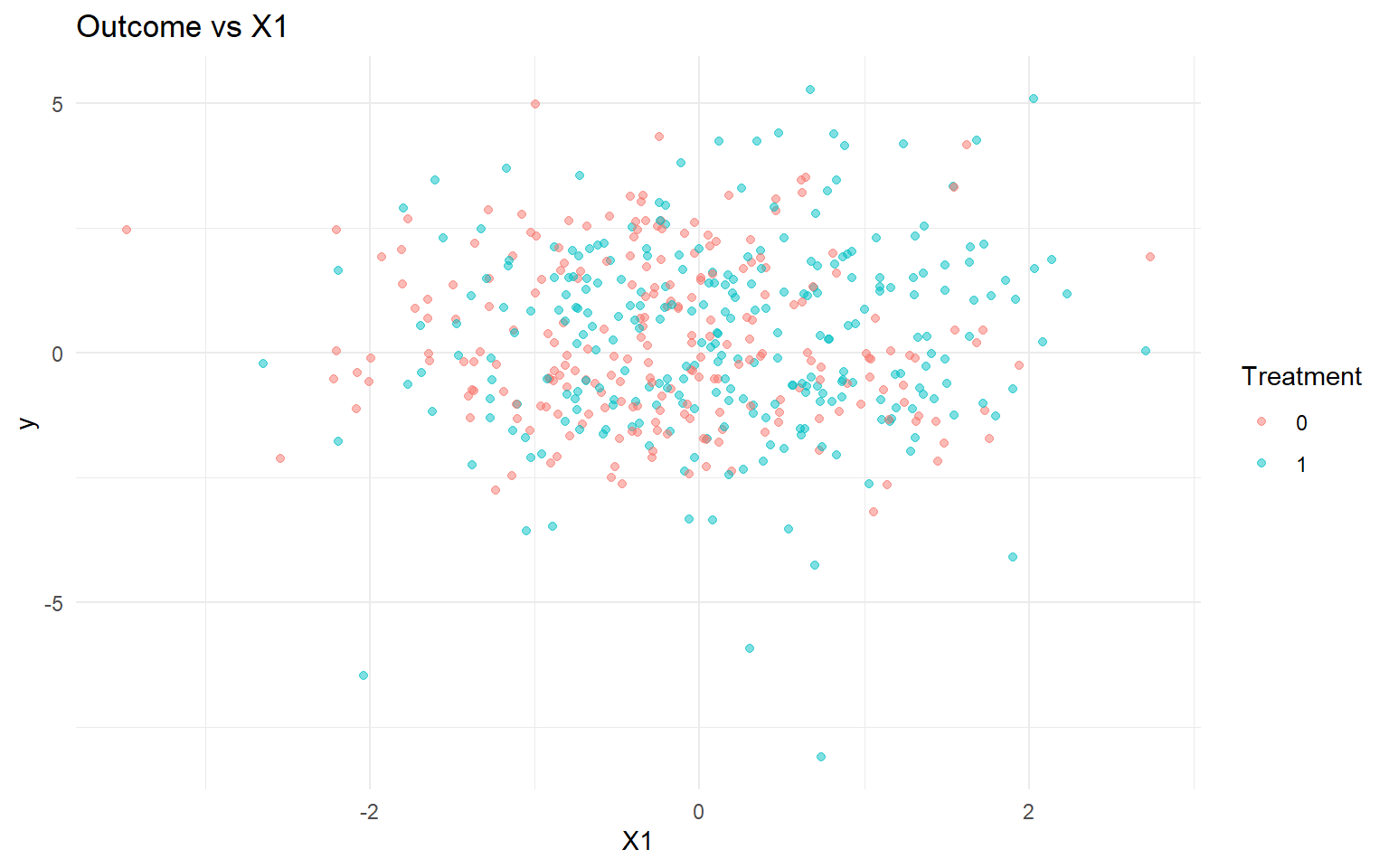
<- data.frame (y = y, A = as.factor (A), x1 = X[, 1 ])<- ggplot (df_causal, aes (x = x1, y = y, color = A)) + geom_point (alpha = 0.5 ) + labs (title = "Outcome vs X1" , x = "X1" , y = "y" , color = "Treatment" ) + theme_minimal ()
Model A: Treated SB, Control CRP (Cauchy)
Code
<- bundle (y = y,treat = A,X = X,kernel = c ("cauchy" , "cauchy" ),backend = c ("sb" , "crp" ),PS = "logit" ,GPD = c (FALSE , FALSE ),components = components_ex13,mcmc_outcome = bulk_mcmc_ex13,mcmc_ps = mcmc_ps_ex13
CausalMixGPD causal bundle
PS model: Bayesian logit (A | X)
Field Treated Control
Backend Stick-Breaking Process Chinese Restaurant Process
Kernel cauchy cauchy
Components 3 3
GPD tail FALSE FALSE
Epsilon 0.025 0.025
Outcome PS included: TRUE
n (control) = 232 | n (treated) = 268
Code
CausalMixGPD causal bundle summary
CausalMixGPD causal bundle
PS model: Bayesian logit (A | X)
Field Treated Control
Backend Stick-Breaking Process Chinese Restaurant Process
Kernel cauchy cauchy
Components 3 3
GPD tail FALSE FALSE
Epsilon 0.025 0.025
Outcome PS included: TRUE
n (control) = 232 | n (treated) = 268
Code
<- load_or_fit (quiet_mcmc (dpmix.causal (bundle_sb_crp, mcmc = causal_run_mcmc))summary (fit_sb_crp)
-- PS fit --
CausalMixGPD PS fit summary
model: logit
n = 500 | predictors = 5
Monitors: beta
Summary table
parameter mean sd q0.025 q0.500 q0.975
beta[1] 0.031 0.186 -0.234 -0.009 0.456
beta[2] 0.463 0.13 0.168 0.524 0.646
beta[3] 0.41 0.209 0.103 0.461 0.82
beta[4] -0.185 0.075 -0.281 -0.117 -0.116
beta[5] 0.235 0.305 -0.42 0.247 0.74
-- Outcome fits --
[control]
MixGPD summary | backend: Chinese Restaurant Process | kernel: Cauchy Distribution | GPD tail: FALSE | epsilon: 0.025
n = 232 | components = 3
Summary
Initial components: 3 | Components after truncation: 3
WAIC: 942.038
lppd: -361.955 | pWAIC: 109.064
Summary table
parameter mean sd q0.025 q0.500 q0.975
weights[1] 0.453 0.063 0.358 0.455 0.599
weights[2] 0.36 0.036 0.297 0.358 0.427
weights[3] 0.187 0.071 0.066 0.183 0.306
alpha 0.449 0.306 0.103 0.368 1.077
scale[1] 0.701 0.149 0.466 0.697 1.046
scale[2] 0.647 0.152 0.382 0.653 0.986
scale[3] 0.622 0.173 0.339 0.626 0.978
beta_location[1,1] -0.27 0.557 -1.096 -0.037 0.39
beta_location[1,2] 0.119 0.347 -0.529 0.119 0.703
beta_location[1,3] -0.242 0.232 -0.688 -0.208 0.02
beta_location[1,4] 0.732 1.441 -0.916 -0.146 2.485
beta_location[2,1] -0.321 0.458 -0.934 -0.375 0.341
beta_location[2,2] -0.071 0.462 -0.873 -0.067 0.713
beta_location[2,3] -0.177 0.214 -0.551 -0.113 0.058
beta_location[2,4] 0.669 1.532 -1.44 1.824 2.329
beta_location[3,1] 0.258 0.309 -0.366 0.289 0.7
beta_location[3,2] -0.478 0.467 -1.265 -0.559 0.713
beta_location[3,3] 0.069 0.32 -0.522 0.058 0.773
beta_location[3,4] -0.584 0.678 -1.429 -0.885 1.572
[treated]
MixGPD summary | backend: Stick-Breaking Process | kernel: Cauchy Distribution | GPD tail: FALSE | epsilon: 0.025
n = 268 | components = 3
Summary
Initial components: 3 | Components after truncation: 3
Summary table
parameter mean sd q0.025 q0.500 q0.975
weights[1] 0.664 0.043 0.593 0.66 0.741
weights[2] 0.201 0.033 0.154 0.203 0.27
weights[3] 0.135 0.025 0.088 0.136 0.182
alpha 1.272 0.641 0.129 1.147 2.63
scale[1] 1.16 0.109 0.971 1.162 1.343
scale[2] 1.199 0.251 0.74 1.19 1.699
scale[3] 1.311 0.38 0.747 1.232 2.277
beta_location[1,1] 0 0 0 0 0
beta_location[1,2] 0 0 0 0 0
beta_location[1,3] 0 0 0 0 0
beta_location[1,4] 0 0 0 0 0
beta_location[2,1] 0 0 0 0 0
beta_location[2,2] 0 0 0 0 0
beta_location[2,3] 0 0 0 0 0
beta_location[2,4] 0 0 0 0 0
beta_location[3,1] 0 0 0 0 0
beta_location[3,2] 0 0 0 0 0
beta_location[3,3] 0 0 0 0 0
beta_location[3,4] 0 0 0 0 0
Code
Posterior mean parameters (causal)
[ps]
Posterior mean parameters
$beta
[1] 0.03087 0.46270 0.40980 -0.18530 0.23550
[treated]
Posterior mean parameters
$alpha
[1] 1.272
$w
[1] 0.6642 0.2008 0.1350
$beta_location
PropScore x1 x2 x3 x4
comp1 0 0 0 0 0
comp2 0 0 0 0 0
comp3 0 0 0 0 0
$scale
[1] 1.160 1.199 1.311
[control]
Posterior mean parameters
$alpha
[1] 0.4488
$w
[1] 0.4528 0.3599 0.1873
$beta_location
PropScore x1 x2 x3 x4
comp1 0.1207 -0.2704 0.11890 -0.24190 0.7316
comp2 0.1856 -0.3205 -0.07111 -0.17740 0.6692
comp3 -0.9230 0.2583 -0.47790 0.06883 -0.5837
$scale
[1] 0.7009 0.6470 0.6218
Code
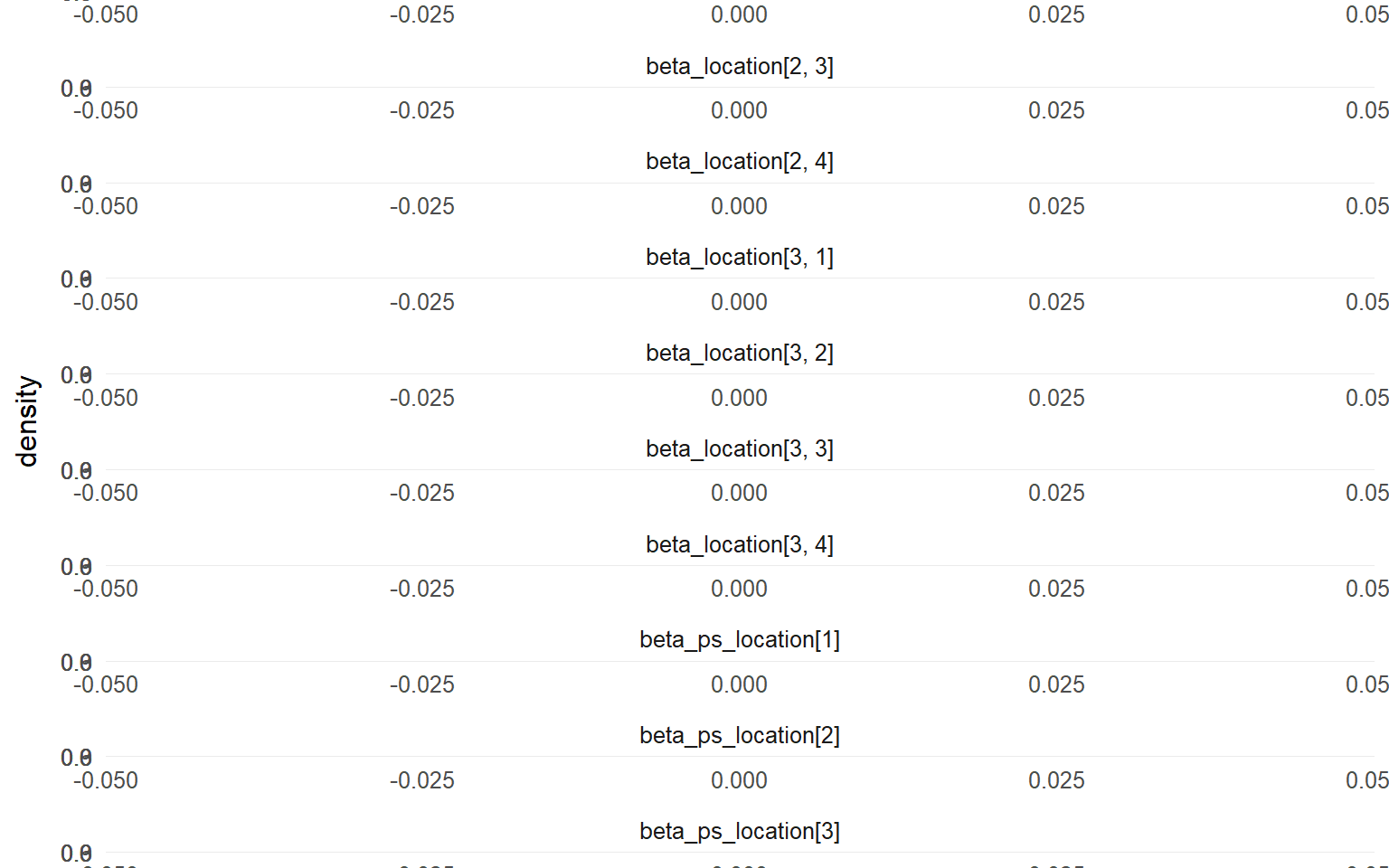
plot (fit_sb_crp, family = plot_family_sb_crp_ex13)
=== treated ===
=== traceplot ===
=== control ===
=== traceplot ===
Code
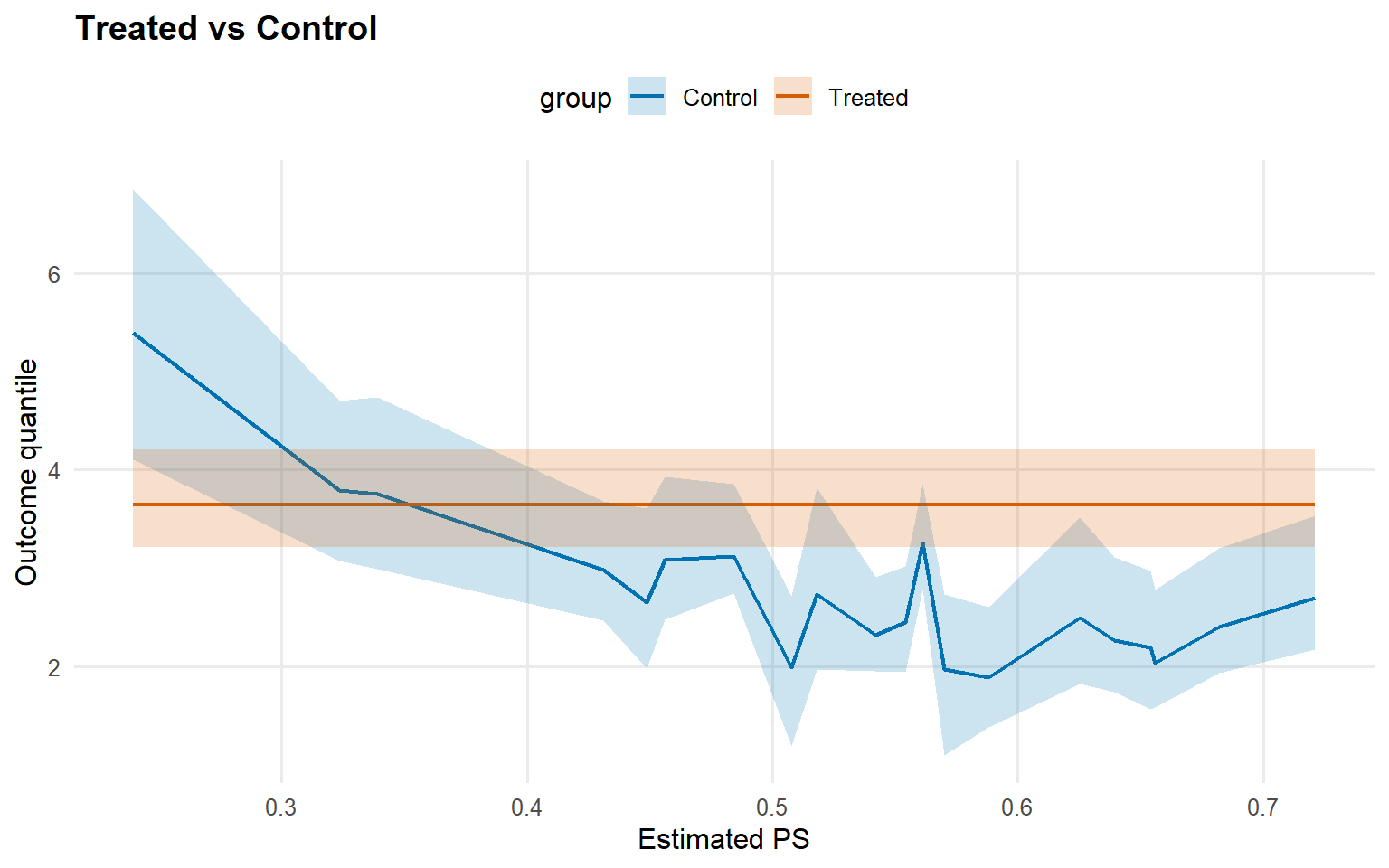
<- predict (newdata = x_eval,type = "quantile" ,p = 0.5 ,interval = "credible" ,workers = predict_workers_ex13plot (pred_median_sb_crp)
Code
<- predict (fit_sb_crp, newdata = x_eval, type = "quantile" ,p = 0.9 , interval = "credible" , workers = predict_workers_ex13)plot (pred_q_sb_crp)
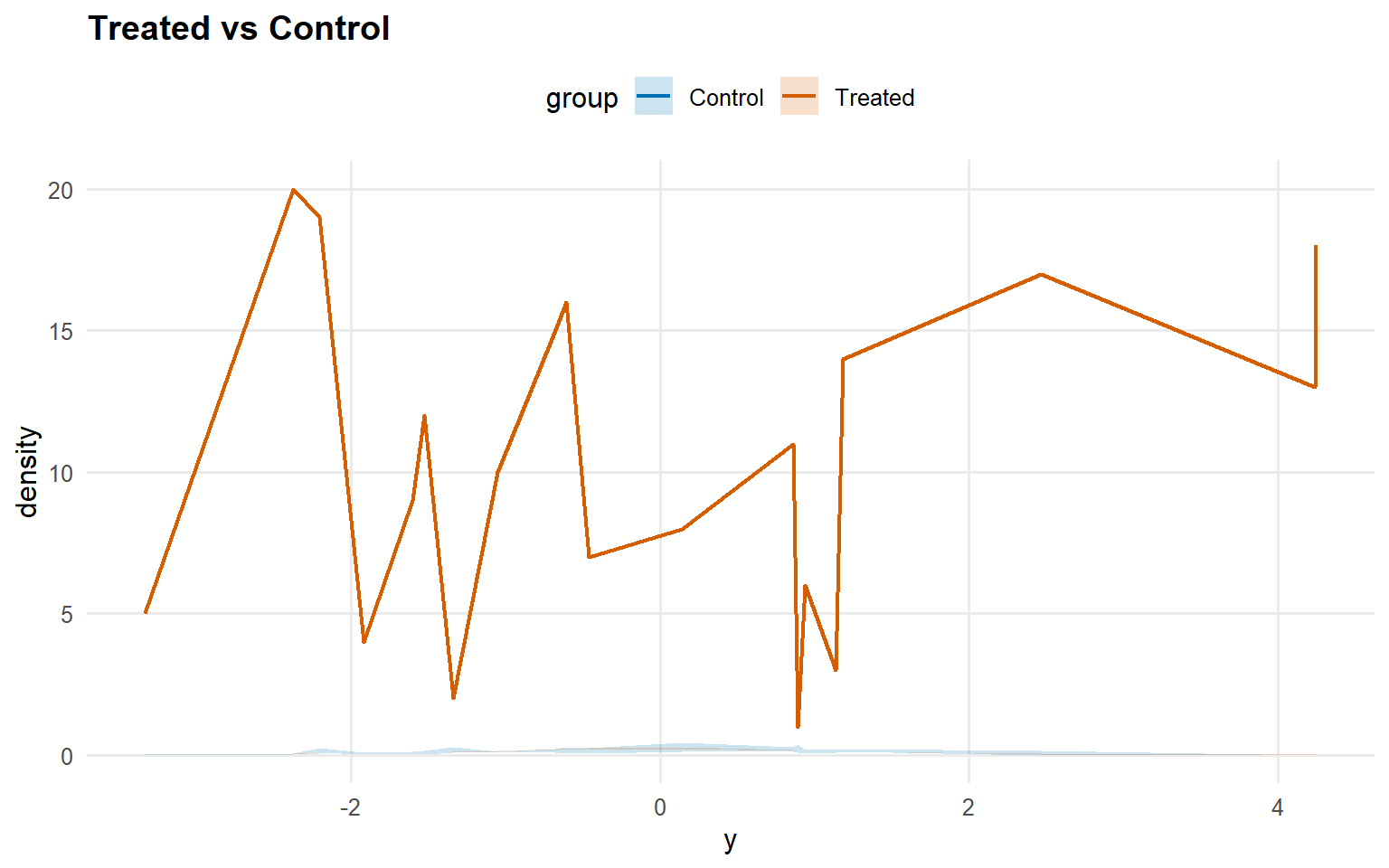
Code
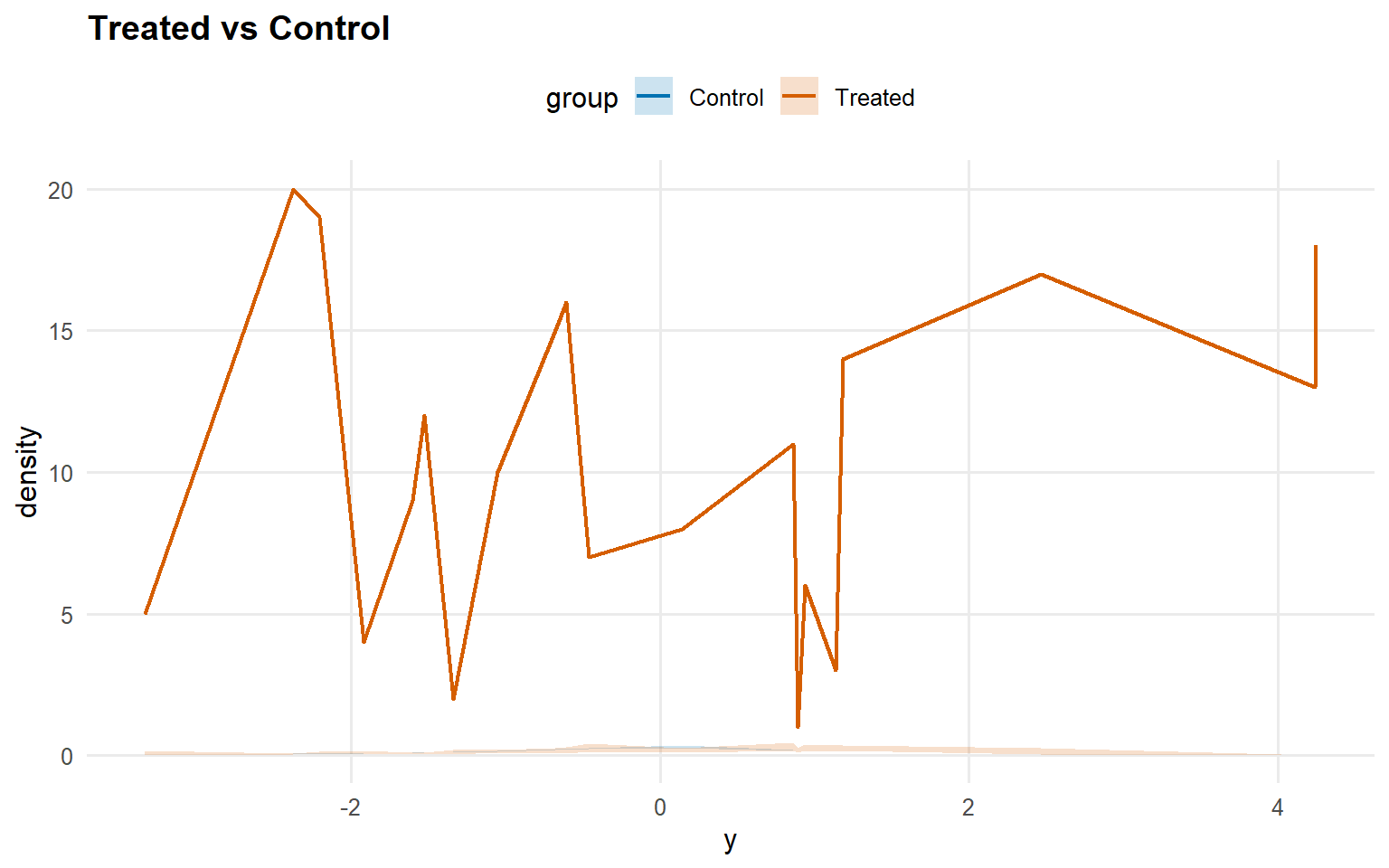
<- predict (fit_sb_crp, newdata = x_eval, y = y_eval,type = "density" , interval = "credible" , workers = predict_workers_ex13)plot (pred_d_sb_crp)
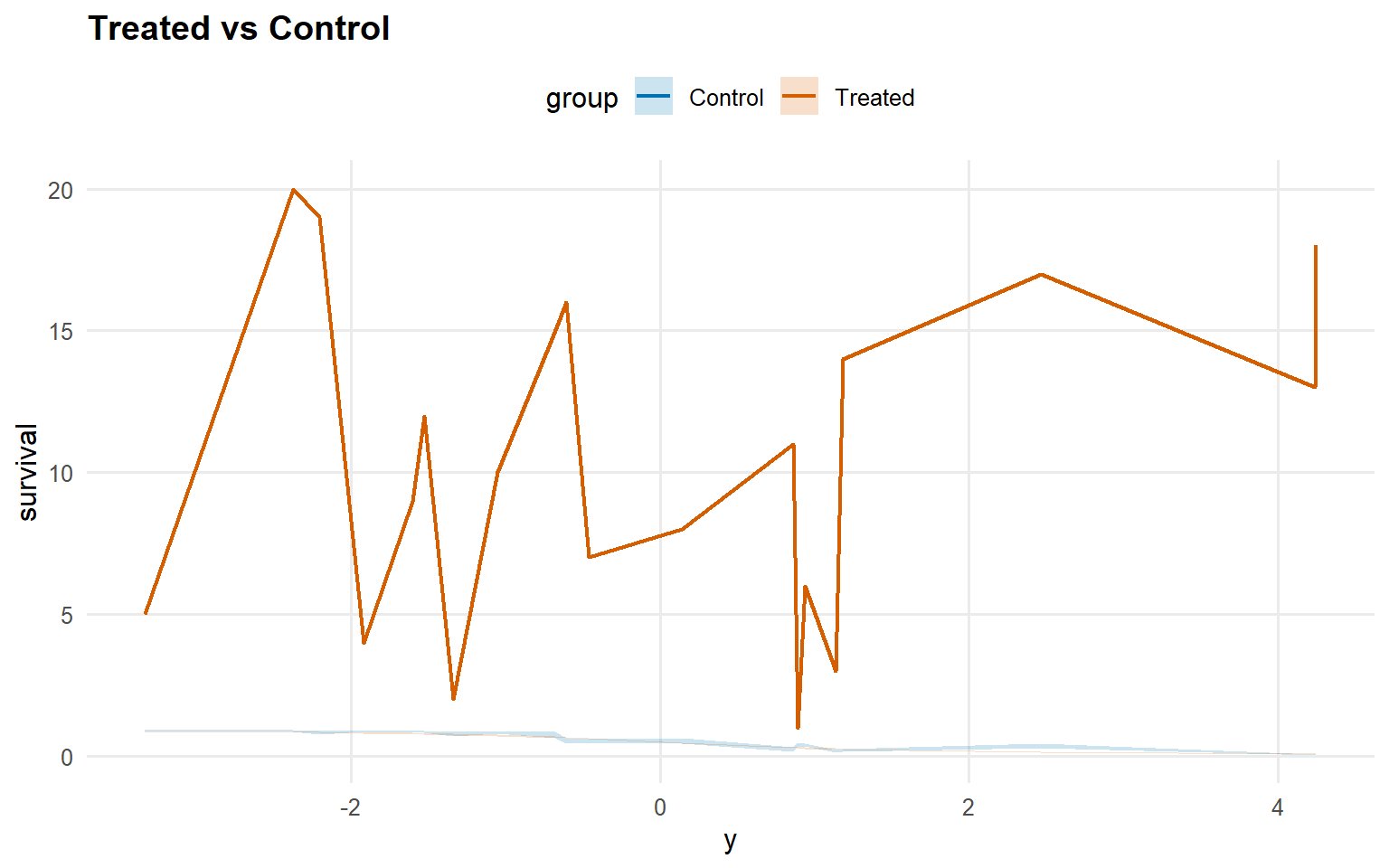
Code
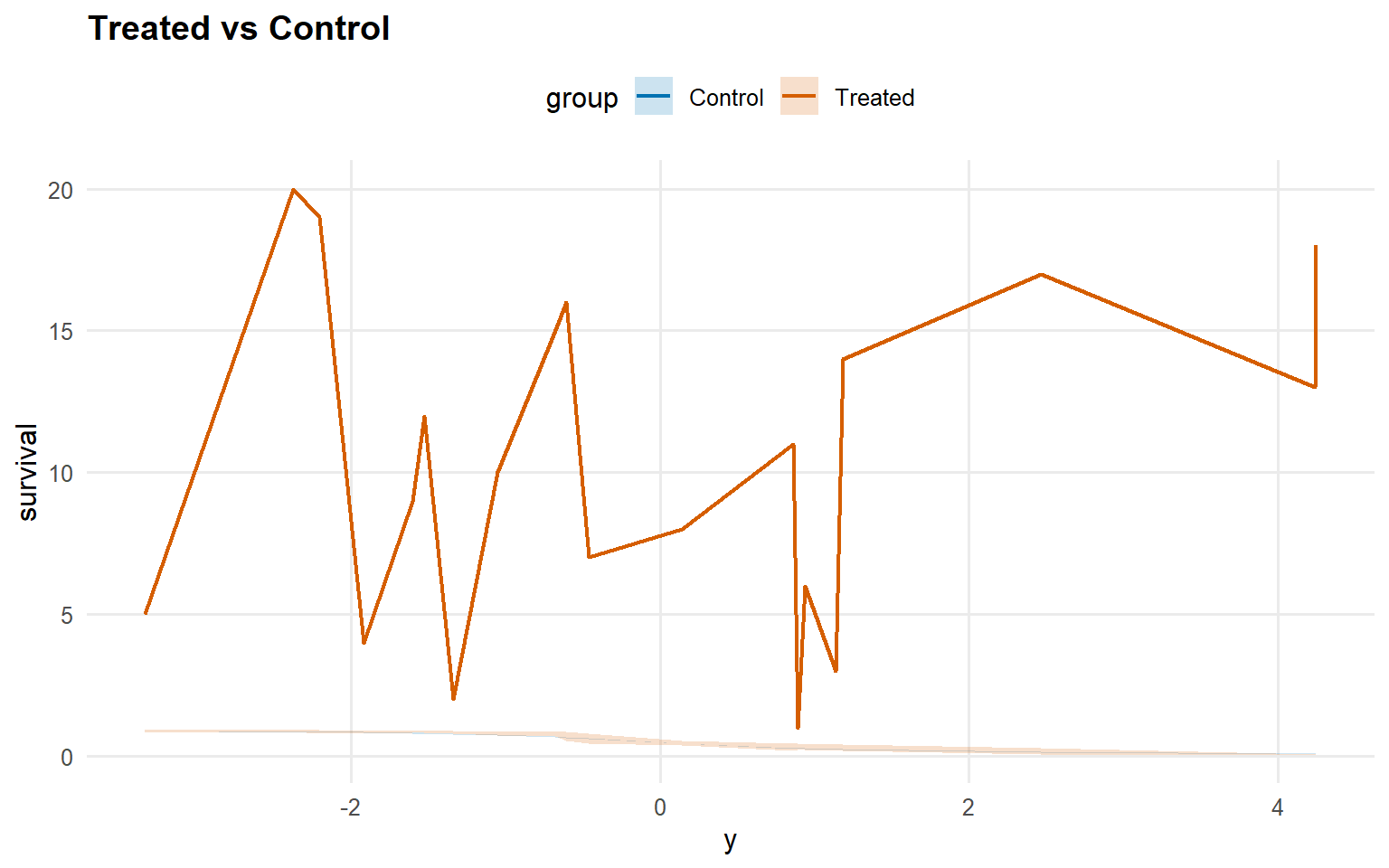
<- predict (fit_sb_crp, newdata = x_eval, y = y_eval,type = "survival" , interval = "credible" , workers = predict_workers_ex13)plot (pred_surv_sb_crp)
Code
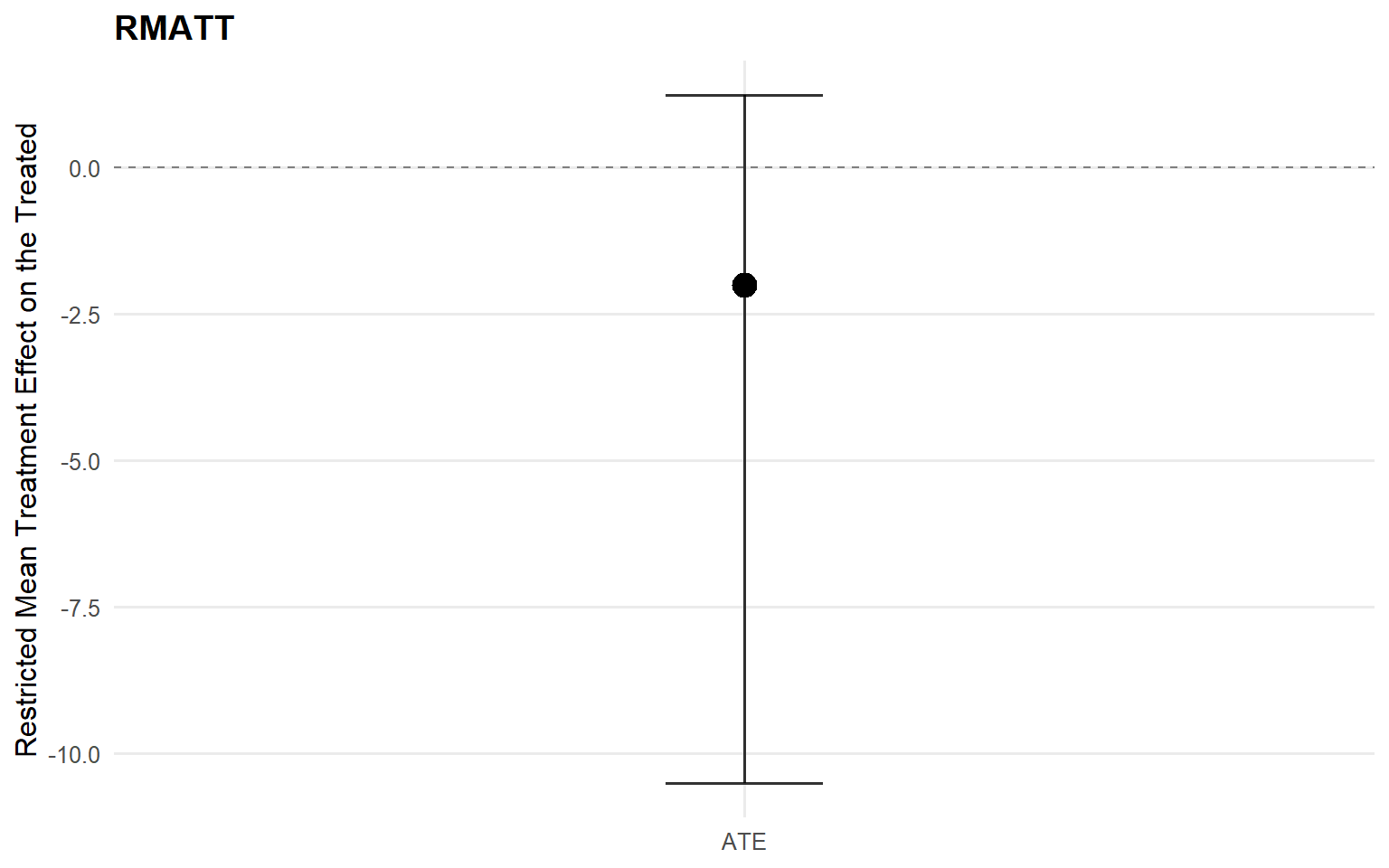
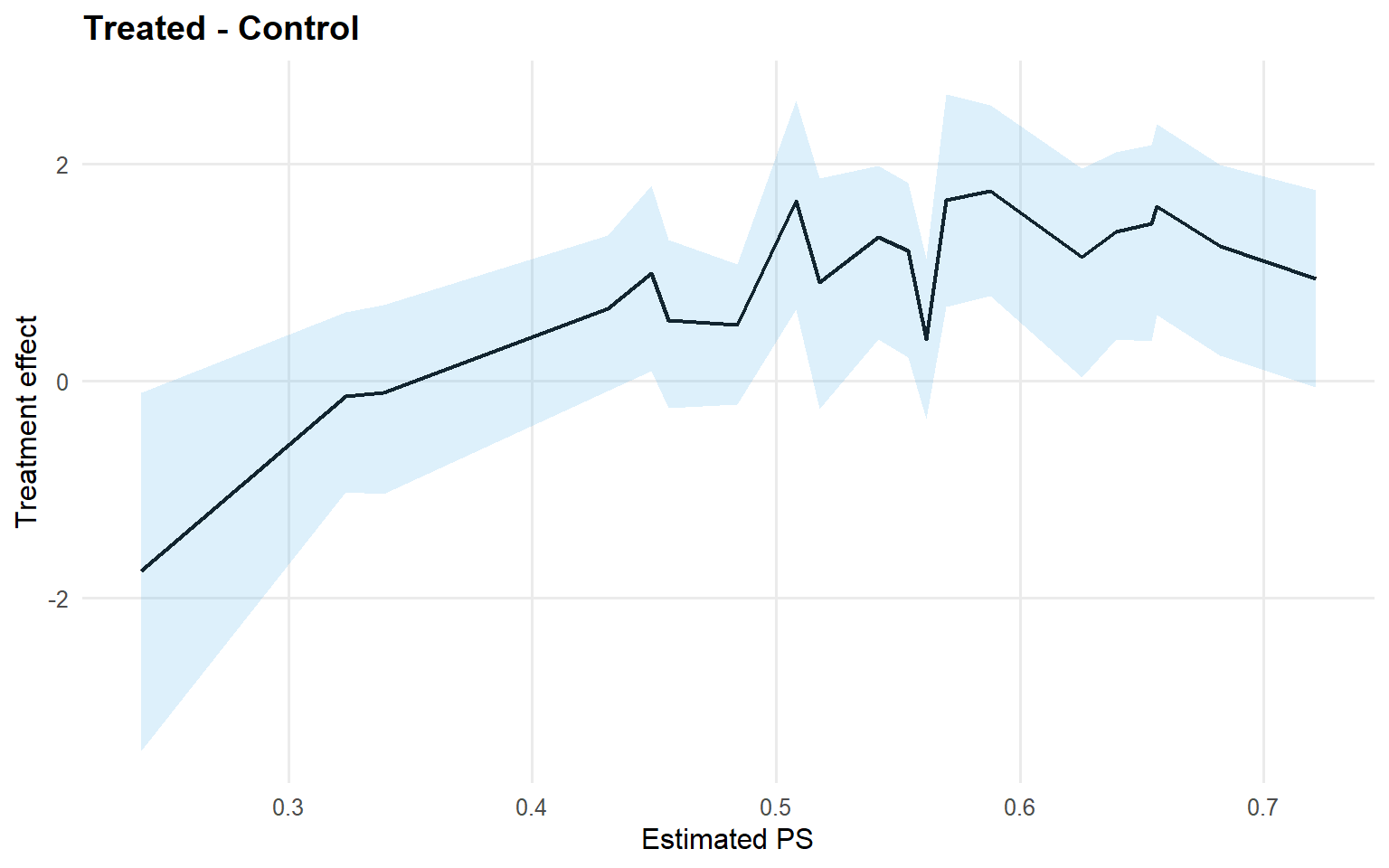
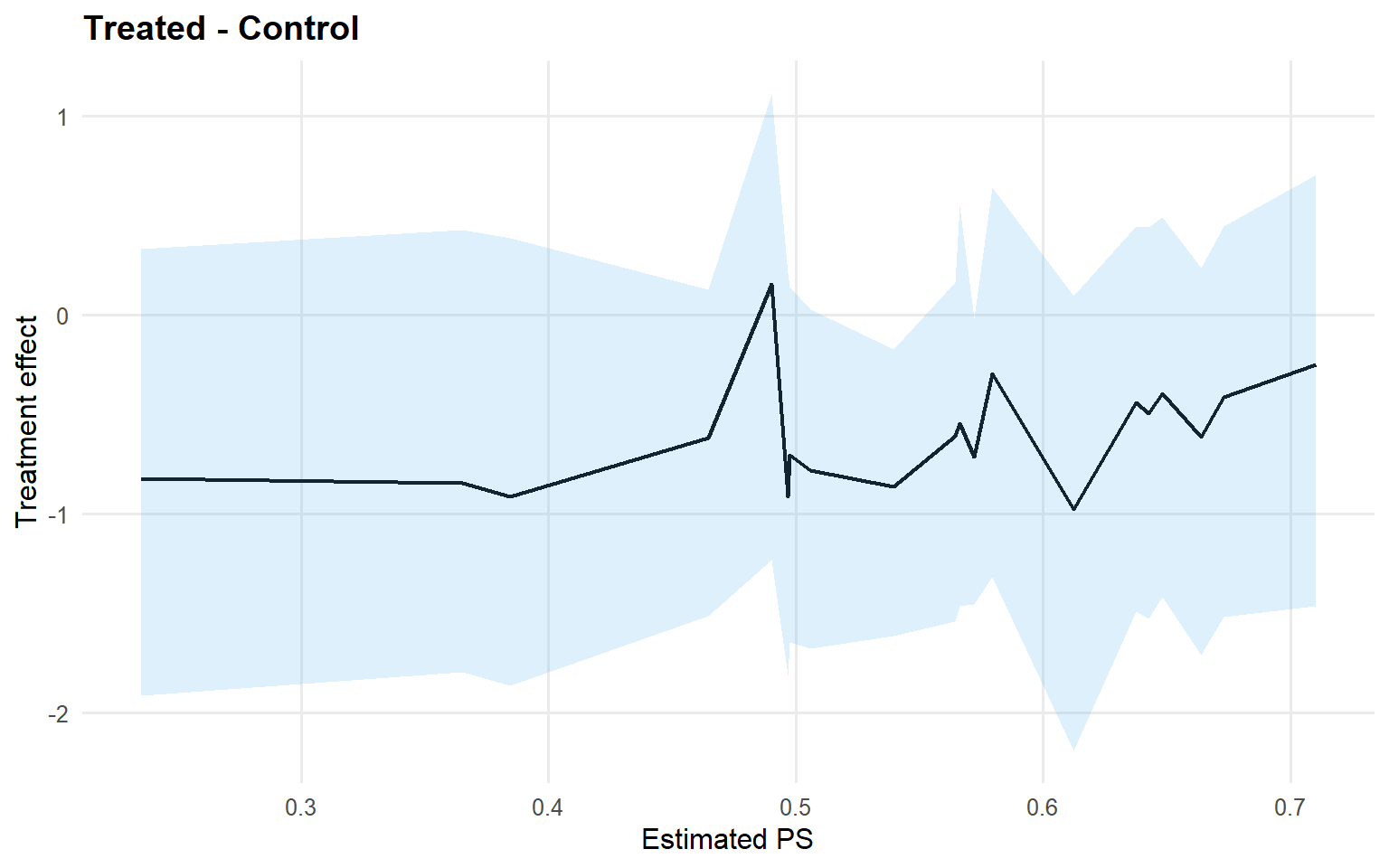
<- att (type = "rmean" ,cutoff = rmean_cutoff_ex13,interval = "credible" ,nsim_mean = nsim_mean_ex13plot (att_sb_crp)
Code
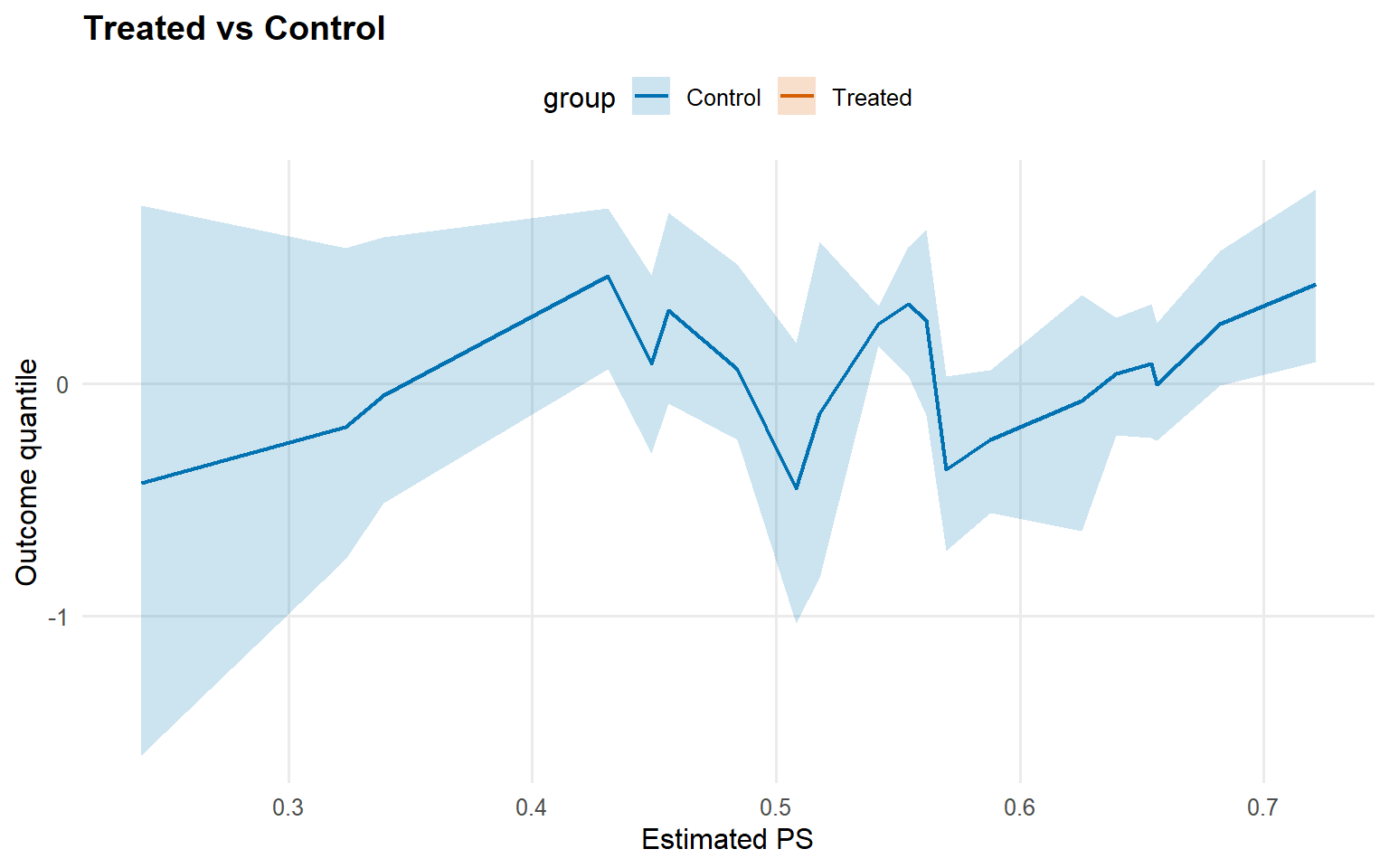
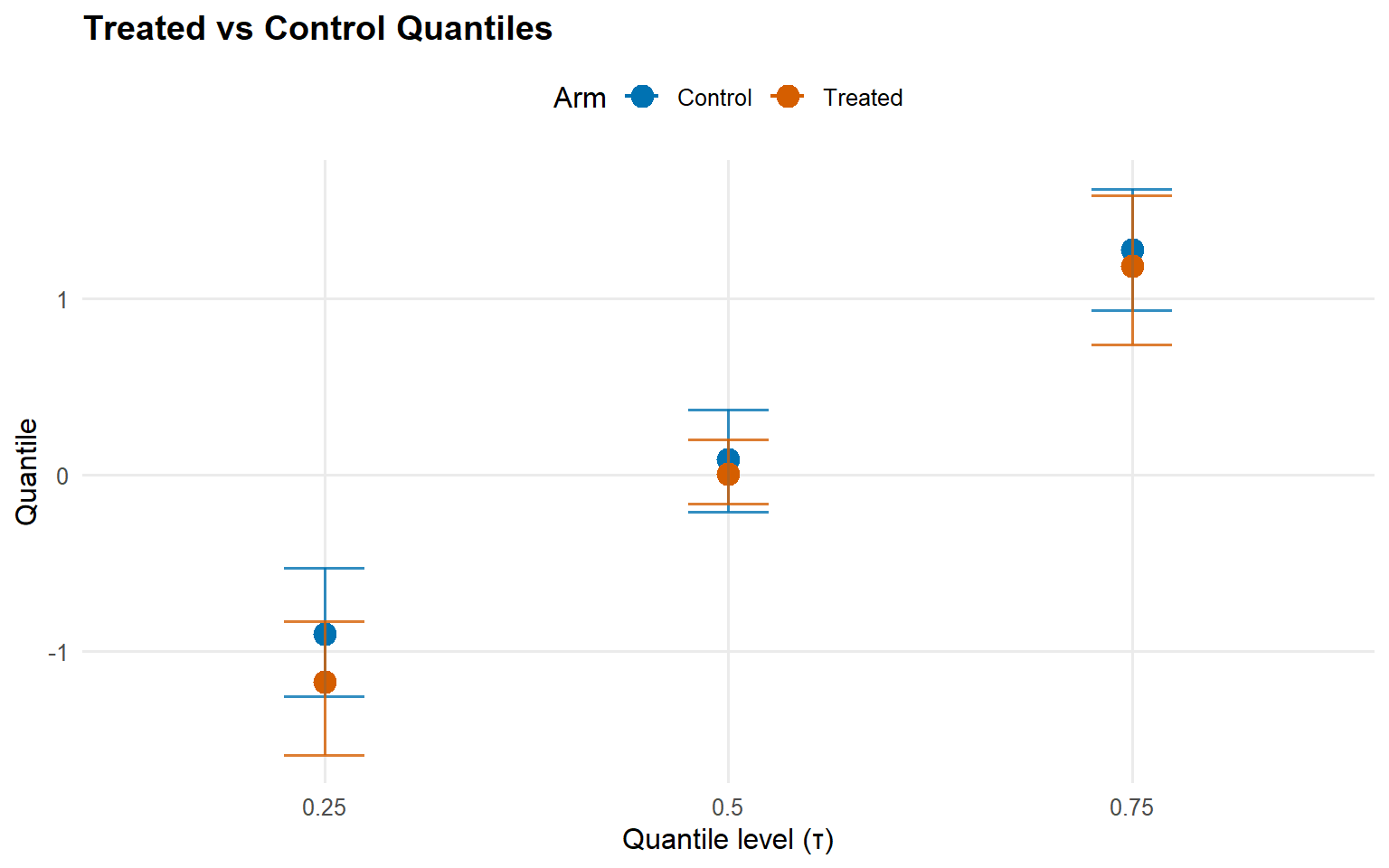
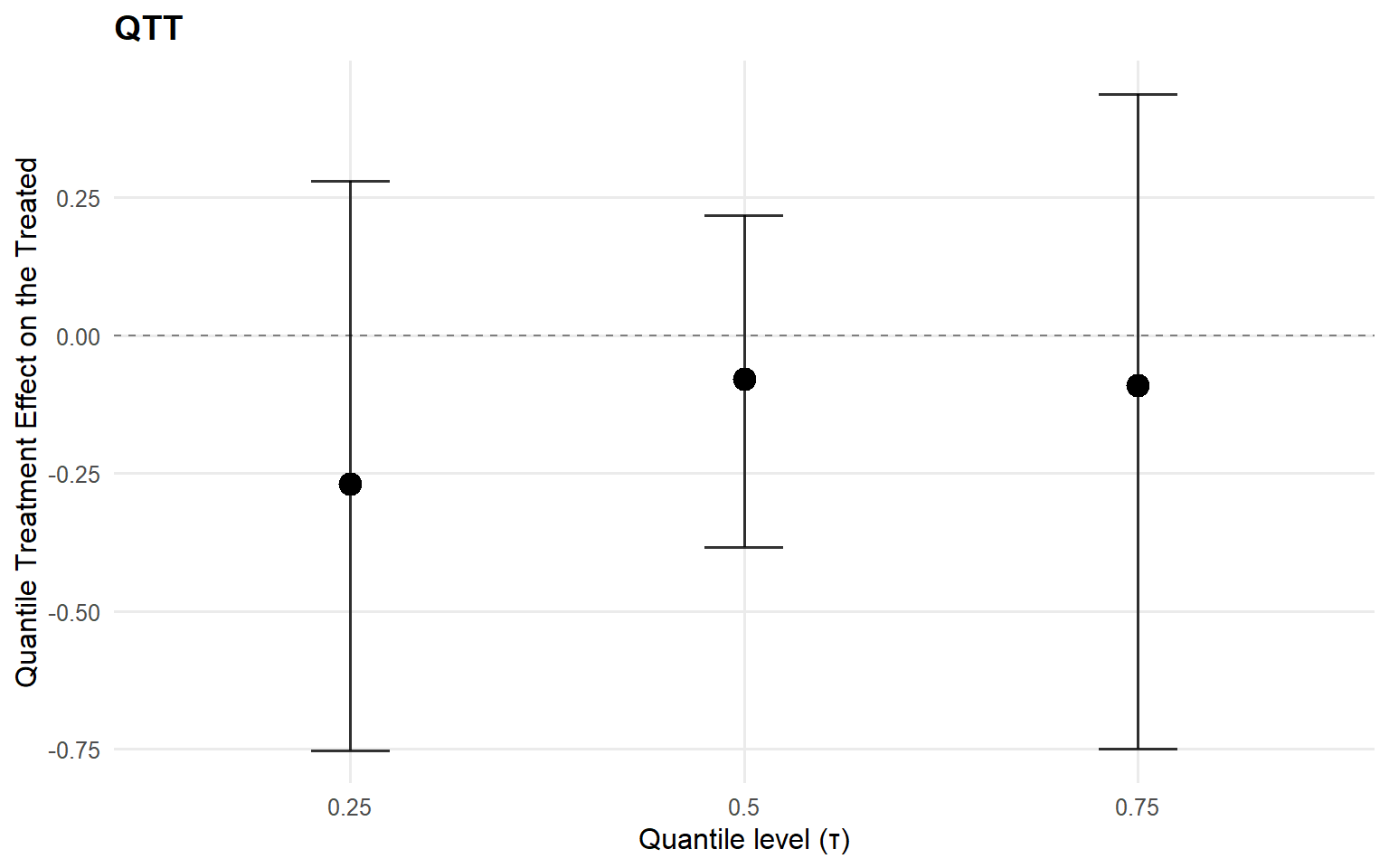
<- qtt (fit_sb_crp, probs = c (0.25 , 0.5 , 0.75 ), interval = "credible" )plot (qtt_sb_crp)
Model B: Treated CRP, Control SB (Cauchy)
Code
<- bundle (y = y,treat = A,X = X,kernel = c ("cauchy" , "cauchy" ),backend = c ("crp" , "sb" ),PS = "logit" ,GPD = c (FALSE , FALSE ),components = components_ex13,mcmc_outcome = bulk_mcmc_ex13_b,mcmc_ps = mcmc_ps_ex13_b
CausalMixGPD causal bundle
PS model: Bayesian logit (A | X)
Field Treated Control
Backend Chinese Restaurant Process Stick-Breaking Process
Kernel cauchy cauchy
Components 3 3
GPD tail FALSE FALSE
Epsilon 0.025 0.025
Outcome PS included: TRUE
n (control) = 232 | n (treated) = 268
Code
CausalMixGPD causal bundle summary
CausalMixGPD causal bundle
PS model: Bayesian logit (A | X)
Field Treated Control
Backend Chinese Restaurant Process Stick-Breaking Process
Kernel cauchy cauchy
Components 3 3
GPD tail FALSE FALSE
Epsilon 0.025 0.025
Outcome PS included: TRUE
n (control) = 232 | n (treated) = 268
Code
<- load_or_fit (quiet_mcmc (dpmix.causal (bundle_crp_sb, mcmc = causal_run_mcmc))summary (fit_crp_sb)
-- PS fit --
CausalMixGPD PS fit summary
model: logit
n = 500 | predictors = 5
Monitors: beta
Summary table
parameter mean sd q0.025 q0.500 q0.975
beta[1] 0.059 0.092 -0.056 0.067 0.264
beta[2] 0.458 0.116 0.215 0.456 0.586
beta[3] 0.258 0.166 0.041 0.2 0.544
beta[4] -0.167 0.093 -0.315 -0.128 -0.028
beta[5] 0.26 0.156 -0.142 0.348 0.458
-- Outcome fits --
[control]
MixGPD summary | backend: Stick-Breaking Process | kernel: Cauchy Distribution | GPD tail: FALSE | epsilon: 0.025
n = 232 | components = 3
Summary
Initial components: 3 | Components after truncation: 3
Summary table
parameter mean sd q0.025 q0.500 q0.975
weights[1] 0.76 0.006 0.747 0.762 0.767
weights[2] 0.166 0.016 0.14 0.174 0.185
weights[3] 0.073 0.018 0.053 0.061 0.098
alpha 0.731 0.343 0.152 0.683 1.473
scale[1] 1.078 0.109 0.87 1.093 1.245
scale[2] 1.187 0.413 0.57 1.176 2.408
scale[3] 1.098 0.471 0.526 1.024 2.012
beta_location[1,1] 0 0 0 0 0
beta_location[1,2] 0 0 0 0 0
beta_location[1,3] 0 0 0 0 0
beta_location[1,4] 0 0 0 0 0
beta_location[2,1] 0 0 0 0 0
beta_location[2,2] 0 0 0 0 0
beta_location[2,3] 0 0 0 0 0
beta_location[2,4] 0 0 0 0 0
beta_location[3,1] 0 0 0 0 0
beta_location[3,2] 0 0 0 0 0
beta_location[3,3] 0 0 0 0 0
beta_location[3,4] 0 0 0 0 0
[treated]
MixGPD summary | backend: Chinese Restaurant Process | kernel: Cauchy Distribution | GPD tail: FALSE | epsilon: 0.025
n = 268 | components = 3
Summary
Initial components: 3 | Components after truncation: 2
WAIC: 1160.258
lppd: -448.99 | pWAIC: 131.139
Summary table
parameter mean sd q0.025 q0.500 q0.975
weights[1] 0.515 0.064 0.415 0.513 0.637
weights[2] 0.397 0.058 0.255 0.403 0.481
alpha 0.427 0.33 0.07 0.35 1.496
scale[1] 0.826 0.168 0.541 0.821 1.125
scale[2] 0.69 0.169 0.448 0.647 1.134
beta_location[1,1] 0.118 0.241 -0.309 0.123 0.523
beta_location[1,2] 0.345 0.885 -0.912 0.135 1.71
beta_location[1,3] 0.002 0.439 -0.739 0.1 0.648
beta_location[1,4] 0.525 0.497 -0.418 0.667 1.177
beta_location[2,1] 0.235 0.324 -0.348 0.229 0.713
beta_location[2,2] 0.348 0.975 -1.214 0.042 1.668
beta_location[2,3] -0.14 0.468 -0.739 0.1 0.398
beta_location[2,4] 0.313 0.516 -0.41 0.369 1.195
Code
Posterior mean parameters (causal)
[ps]
Posterior mean parameters
$beta
[1] 0.05896 0.45800 0.25750 -0.16720 0.26020
[treated]
Posterior mean parameters
$alpha
[1] 0.427
$w
[1] 0.5148 0.3970
$beta_location
PropScore x1 x2 x3 x4
comp1 -0.3378 0.1177 0.3445 0.001586 0.5255
comp2 -0.3749 0.2346 0.3484 -0.140100 0.3132
$scale
[1] 0.8260 0.6899
[control]
Posterior mean parameters
$alpha
[1] 0.7308
$w
[1] 0.76030 0.16640 0.07332
$beta_location
PropScore x1 x2 x3 x4
comp1 0 0 0 0 0
comp2 0 0 0 0 0
comp3 0 0 0 0 0
$scale
[1] 1.078 1.187 1.098
Code
plot (fit_crp_sb, family = plot_family_crp_sb_ex13)
=== treated ===
=== density ===
=== control ===
=== density ===
Code
<- predict (newdata = x_eval,type = "quantile" ,p = 0.5 ,interval = "credible" ,workers = predict_workers_ex13plot (pred_median_crp_sb)
Code
<- predict (fit_crp_sb, newdata = x_eval, type = "quantile" ,p = 0.9 , interval = "credible" , workers = predict_workers_ex13)plot (pred_q_crp_sb)
Code
<- predict (fit_crp_sb, newdata = x_eval, y = y_eval,type = "density" , interval = "credible" , workers = predict_workers_ex13)plot (pred_d_crp_sb)
Code
<- predict (fit_crp_sb, newdata = x_eval, y = y_eval,type = "survival" , interval = "credible" , workers = predict_workers_ex13)plot (pred_surv_crp_sb)
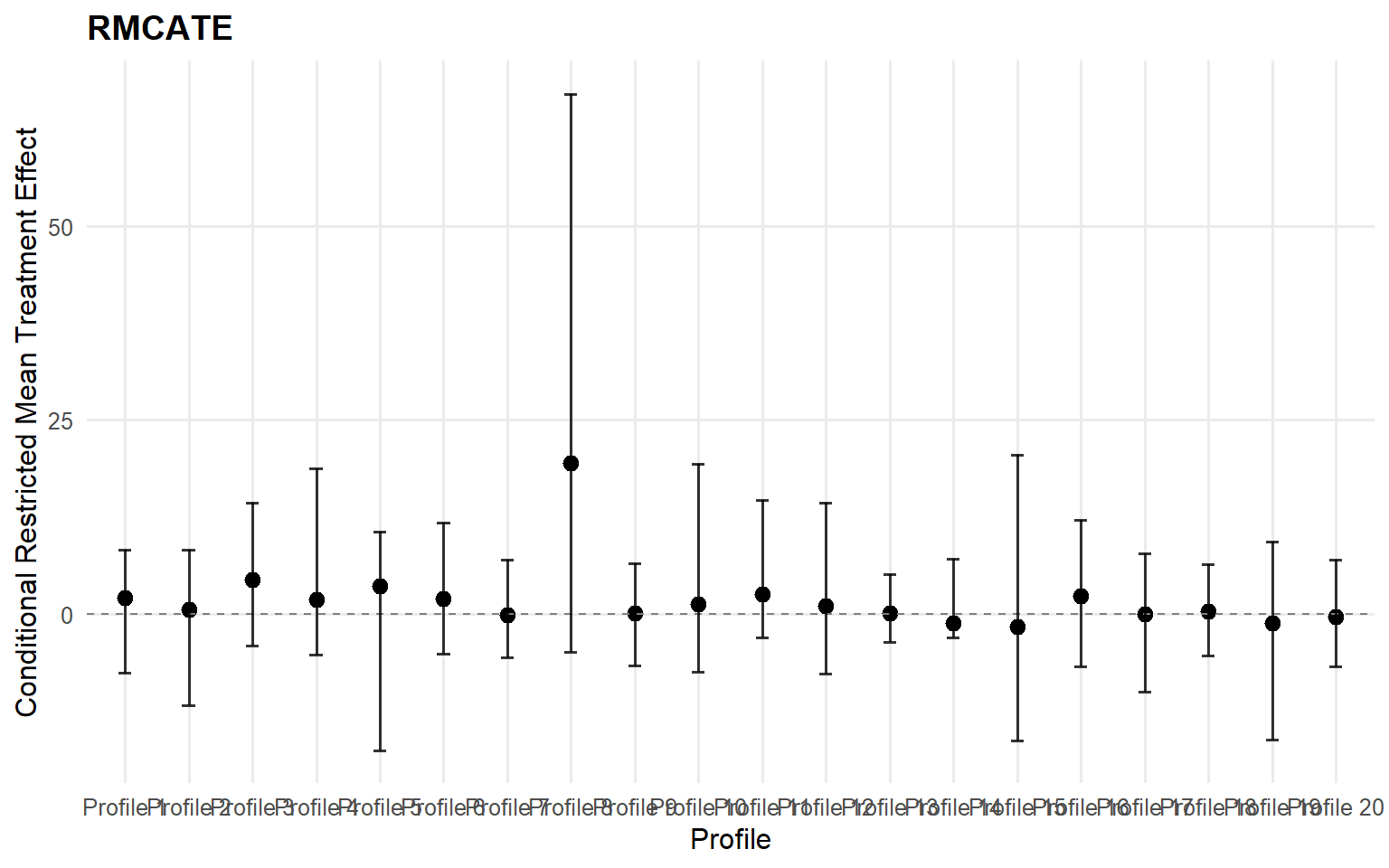
Code
<- cate (type = "rmean" ,cutoff = rmean_cutoff_ex13,newdata = x_eval,interval = "credible" ,nsim_mean = nsim_mean_ex13head (format_cate_table (cate_crp_sb))
id index estimate lower upper
1 1 1 2.0712056 -7.601197 8.232255
2 2 1 0.5246956 -11.766421 8.257291
3 3 1 4.3585056 -4.097305 14.317757
4 4 1 1.8263269 -5.285917 18.786957
5 5 1 3.5740492 -17.630042 10.558621
6 6 1 1.9830236 -5.209355 11.786967
Code
Code
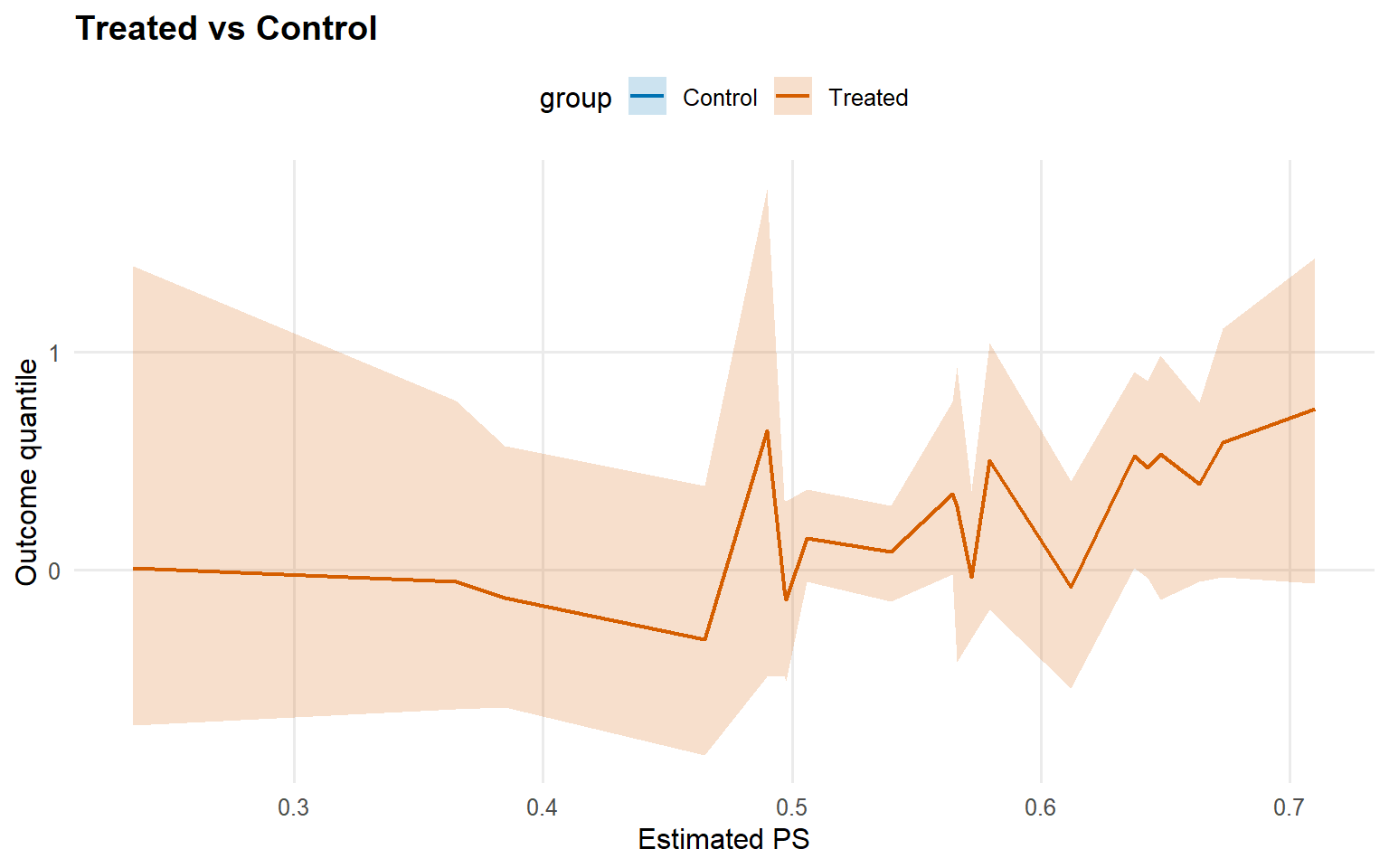
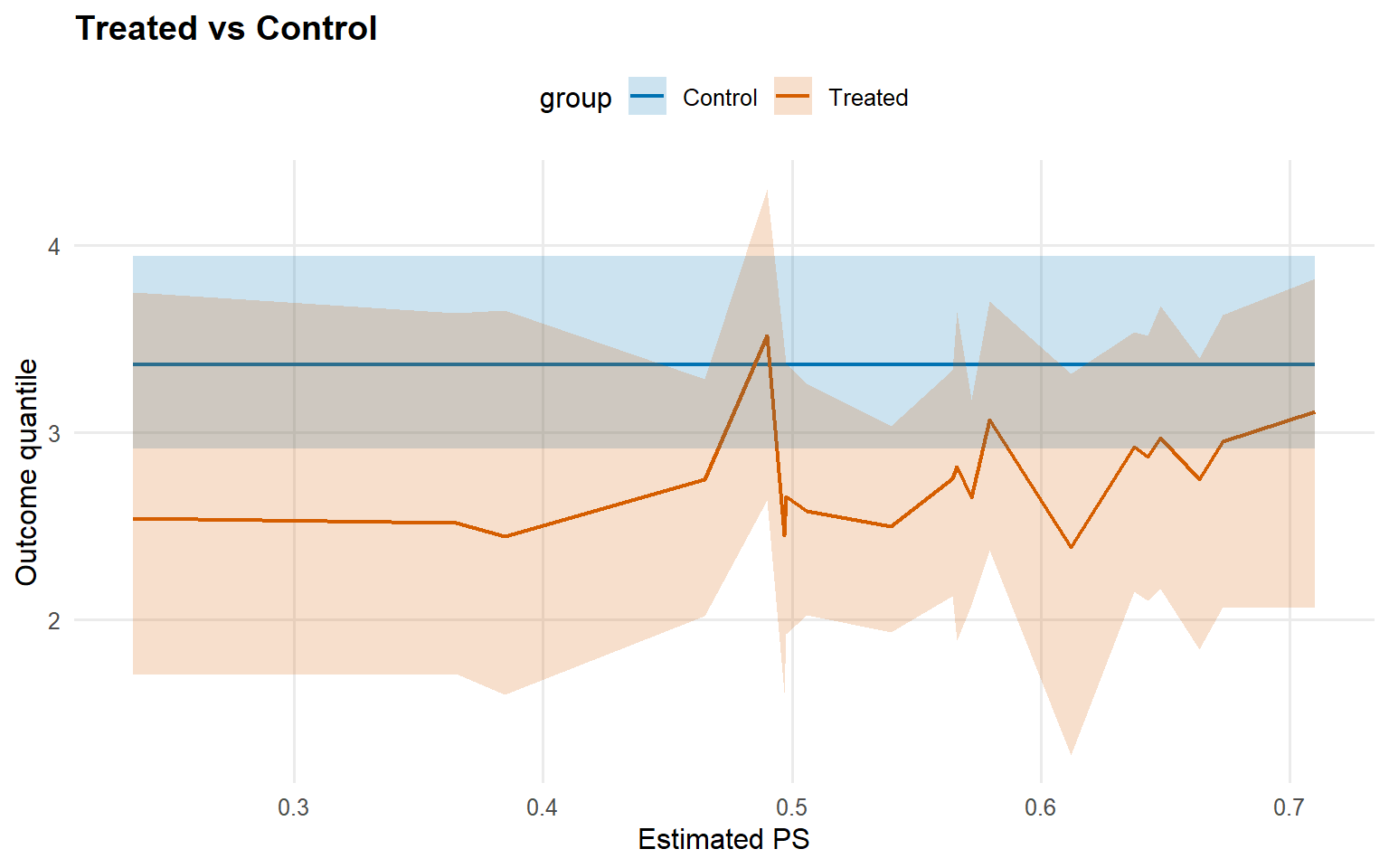
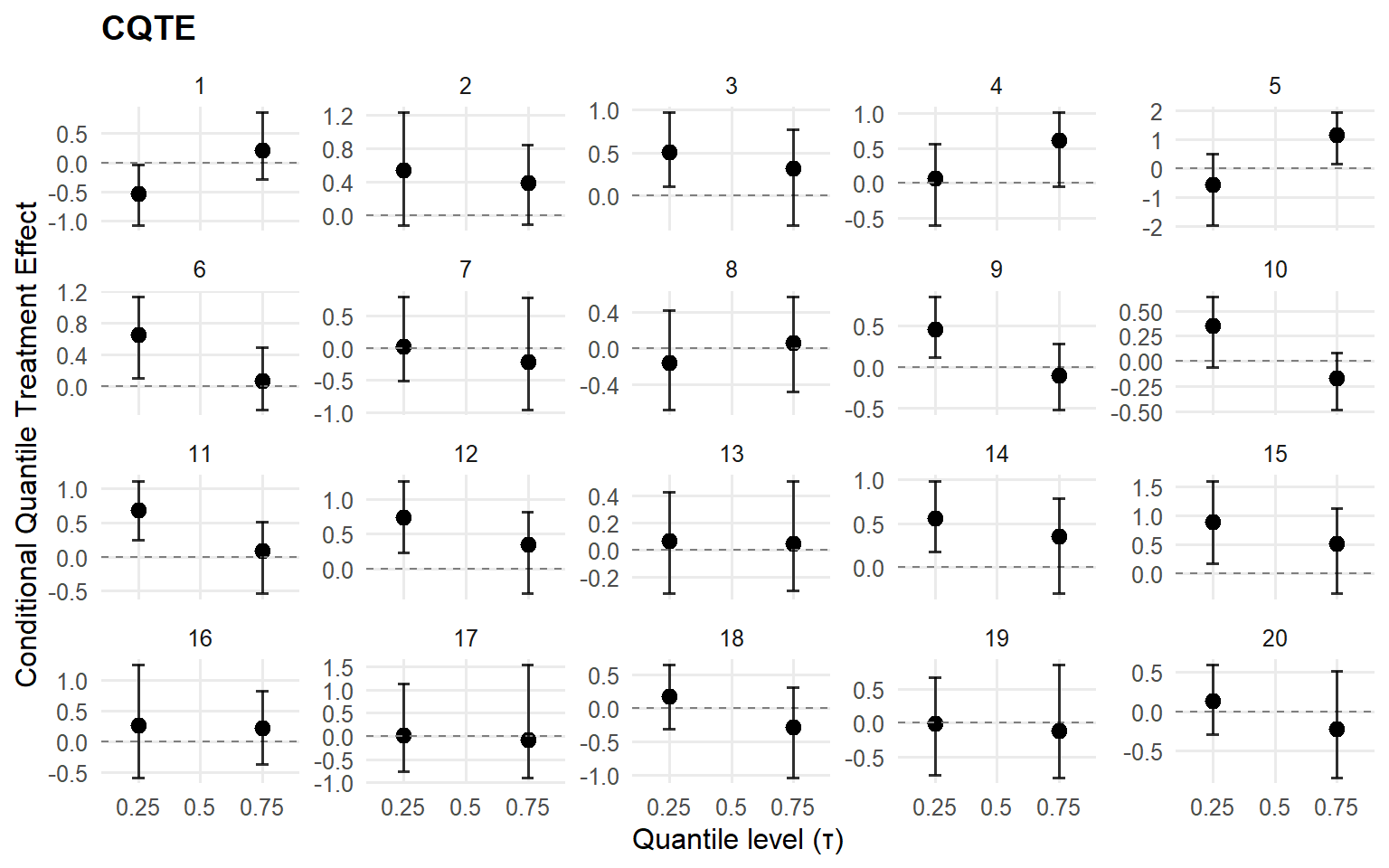
<- cqte (fit_crp_sb, probs = c (0.25 , 0.5 , 0.75 ),newdata = x_eval, interval = "credible" )head (format_cqte_table (cqte_crp_sb))
id index estimate lower upper
1 1 1 -0.5323511 -1.0682313 -0.04068125
2 1 2 NaN NA NA
3 1 3 0.2101863 -0.2865770 0.85255201
4 2 1 0.5353392 -0.1215984 1.22619823
5 2 2 NaN NA NA
6 2 3 0.3822083 -0.1154588 0.83868138
Code
Prereqs
Required packages and data for this page are listed in the setup chunks above.
Outputs
This page renders model fits, diagnostics, and summary artifacts generated by package APIs.
Interpretation
Canonical concept page: 03 Causal Inference Objects
Treat this page as an application/example view and use the canonical page for core definitions.
Next
Continue to the linked canonical concept page, then return for implementation-specific details.