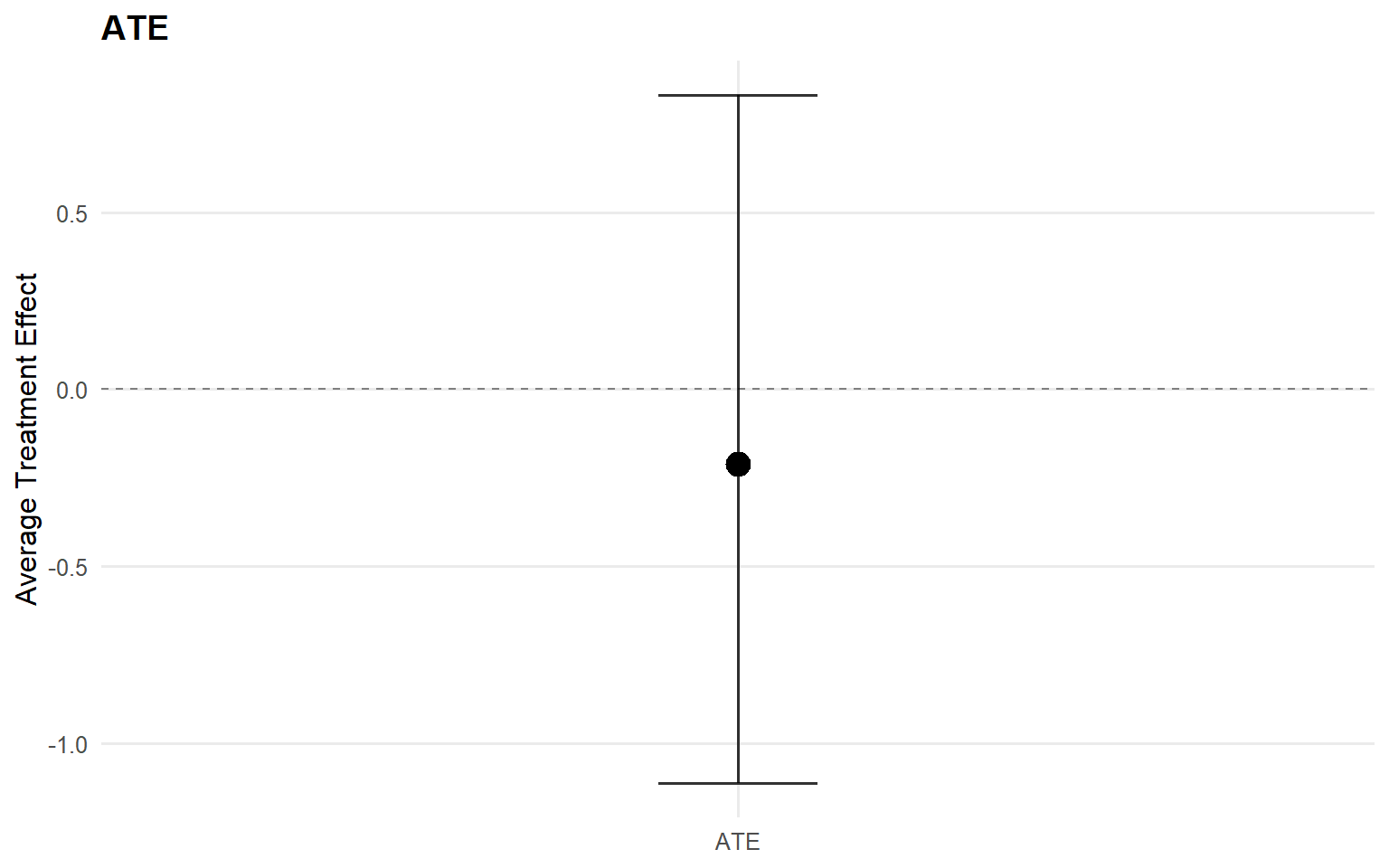Website workflow note. This page reflects the current exported API and recommended wrapper-first usage. Last updated: 2026-02-19 .
For the full package narrative, see the main package vignettes (basic, unconditional, conditional, and causal).
Causal Inference: Mixed Backends with GPD - Amoroso Kernel
This vignette fits two mixed-backend causal models with GPD tails and an Amoroso kernel:
Model A: Treated = SB, Control = CRP
Model B: Treated = CRP, Control = SB
We shift y to keep outcomes in a stable positive range.
What you’ll learn
How to fit mixed-backend causal models when outcomes are shifted positive and modeled with Amoroso bulk + GPD tails .
How tail augmentation influences survival/quantile predictions and downstream effect summaries.
How to keep model comparisons interpretable by reusing the same causal S3 workflow. ## When to use this template
Outcomes are (or can be made) positive and you need a bulk family like Amoroso plus an explicit extreme-tail model.
You want a stress-test style causal analysis where extremes matter and may differ by treatment arm. ## Key limitation (important)
Shifting outcomes changes the scale: interpret effects on the shifted scale (or transform back carefully if you need original-scale statements). ## Next steps
Compare against the bulk-only mixed-backend setup to isolate the incremental impact of GPD = TRUE.
Data Setup
Code
data ("causal_alt_real500_p4_k2" )<- causal_alt_real500_p4_k2$ y<- causal_alt_real500_p4_k2$ A<- as.matrix (causal_alt_real500_p4_k2$ X)<- abs (min (y_raw)) + 0.1 <- y_raw + shift<- tibble (statistic = c ("N" , "Mean" , "SD" , "Min" , "Max" ),value = c (length (y), mean (y), sd (y), min (y), max (y))
# A tibble: 5 × 2
statistic value
<chr> <dbl>
1 N 500
2 Mean 8.46
3 SD 1.76
4 Min 0.1000
5 Max 13.5
Code
<- X[seq_len (if (isTRUE (FAST)) 20 L else 40 L), , drop = FALSE ]<- y[seq_len (if (isTRUE (FAST)) 20 L else 40 L)]<- as.numeric (stats:: quantile (y, 0.8 , names = FALSE ))
Code
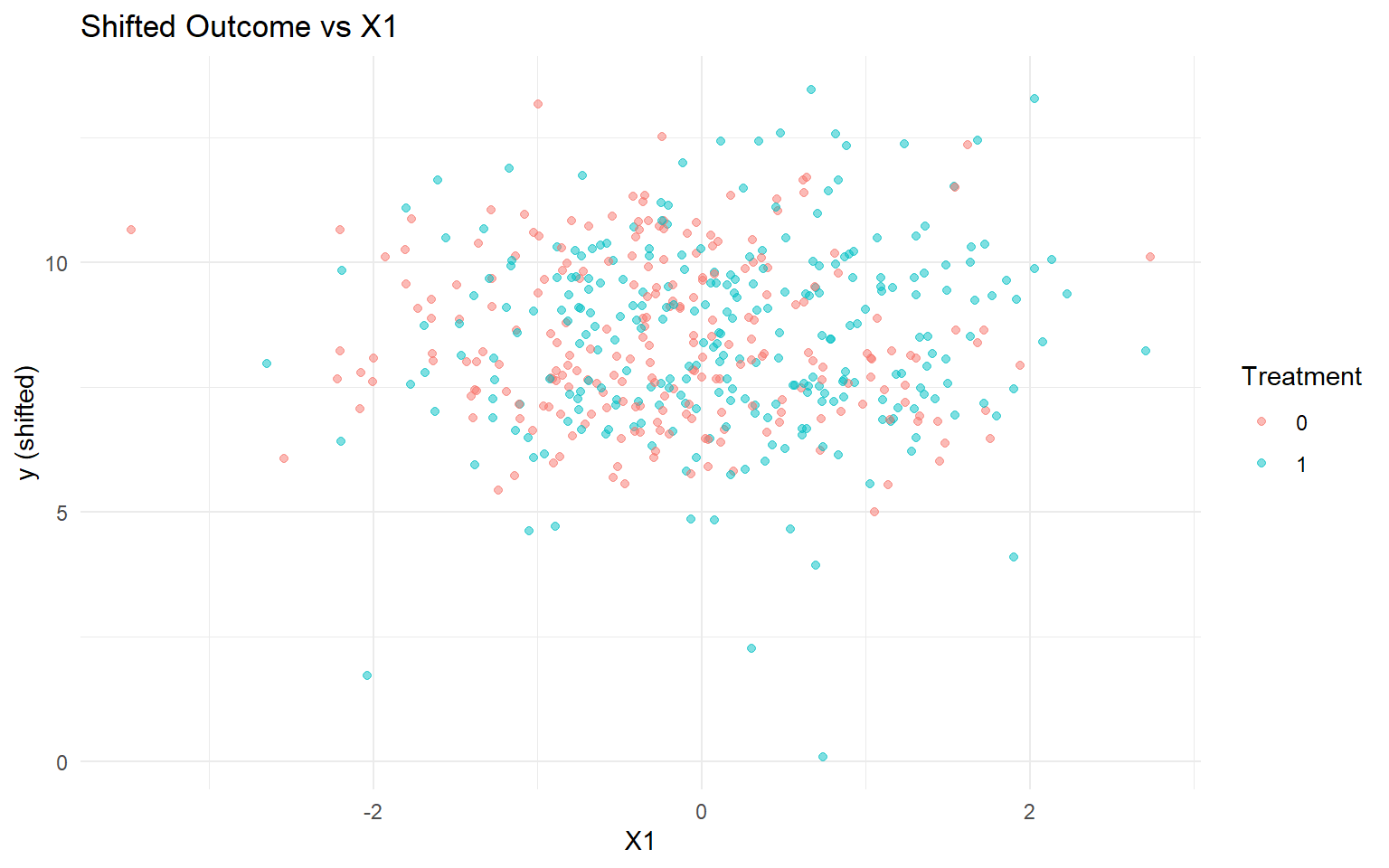
<- data.frame (y = y, A = as.factor (A), x1 = X[, 1 ])<- ggplot (df_causal, aes (x = x1, y = y, color = A)) + geom_point (alpha = 0.5 ) + labs (title = "Shifted Outcome vs X1" , x = "X1" , y = "y (shifted)" , color = "Treatment" ) + theme_minimal ()
Model A: Treated SB, Control CRP (Amoroso + GPD)
Code
<- list (gpd = list (threshold = list (mode = "dist" ,dist = "lognormal" ,# Use max() to guard against log(0) = -Inf by enforcing a positive lower bound. args = list (meanlog = log (max (u_threshold, .Machine$ double.eps)), sdlog = 0.25 )<- bundle (y = y,treat = A,X = X,kernel = c ("amoroso" , "amoroso" ),backend = c ("sb" , "crp" ),PS = "logit" ,GPD = c (TRUE , TRUE ),components = components_ex14,param_specs = param_specs_gpd,mcmc_outcome = gpd_mcmc_ex14,mcmc_ps = mcmc_ps_ex14
CausalMixGPD causal bundle
PS model: Bayesian logit (A | X)
Field Treated Control
Backend Stick-Breaking Process Chinese Restaurant Process
Kernel amoroso amoroso
Components 3 3
GPD tail TRUE TRUE
Epsilon 0.025 0.025
Outcome PS included: TRUE
n (control) = 232 | n (treated) = 268
Code
CausalMixGPD causal bundle summary
CausalMixGPD causal bundle
PS model: Bayesian logit (A | X)
Field Treated Control
Backend Stick-Breaking Process Chinese Restaurant Process
Kernel amoroso amoroso
Components 3 3
GPD tail TRUE TRUE
Epsilon 0.025 0.025
Outcome PS included: TRUE
n (control) = 232 | n (treated) = 268
Code
<- load_or_fit (quiet_mcmc (dpmgpd.causal (bundle_sb_crp, mcmc = causal_run_mcmc))summary (fit_sb_crp)
-- PS fit --
CausalMixGPD PS fit summary
model: logit
n = 500 | predictors = 5
Monitors: beta
Summary table
parameter mean sd q0.025 q0.500 q0.975
beta[1] 0.11 0.25 -0.271 -0.091 0.456
beta[2] 0.474 0.1 0.253 0.509 0.576
beta[3] 0.394 0.186 0.135 0.395 0.656
beta[4] -0.15 0.048 -0.277 -0.117 -0.117
beta[5] 0.101 0.365 -0.42 0.134 0.74
-- Outcome fits --
[control]
MixGPD summary | backend: Chinese Restaurant Process | kernel: Amoroso Distribution | GPD tail: TRUE | epsilon: 0.025
n = 232 | components = 3
Summary
Initial components: 3 | Components after truncation: 1
WAIC: 995.699
lppd: -485.221 | pWAIC: 12.629
Summary table
parameter mean sd q0.025 q0.500 q0.975
weights[1] 1 0 1 1 1
alpha 0.192 0.186 0.008 0.123 0.712
beta_tail_scale[1] 0.089 0.082 -0.055 0.074 0.265
beta_tail_scale[2] 0.059 0.119 -0.216 0.066 0.272
beta_tail_scale[3] 0.023 0.087 -0.145 0.032 0.159
beta_tail_scale[4] 0.365 0.159 0.084 0.376 0.681
tail_shape -0.234 0.047 -0.28 -0.242 -0.081
threshold 9.492 0.014 9.454 9.488 9.504
shape1[1] 5.2 0.283 4.814 5.245 5.585
shape2[1] 0.705 0.005 0.705 0.705 0.705
beta_loc[1,1] -0.943 0.16 -1.328 -0.943 -0.721
beta_loc[1,2] 0.119 0.389 -0.708 0.213 0.559
beta_loc[1,3] 1.118 0.164 0.719 1.146 1.323
beta_loc[1,4] 4.761 1.003 2.042 5.202 5.328
beta_scale[1,1] -0.146 0.168 -0.421 -0.159 0.126
beta_scale[1,2] -0.278 0.117 -0.489 -0.269 -0.081
beta_scale[1,3] -0.221 0.083 -0.331 -0.246 0.002
beta_scale[1,4] -1.082 0.28 -1.437 -1.158 -0.398
[treated]
MixGPD summary | backend: Stick-Breaking Process | kernel: Amoroso Distribution | GPD tail: TRUE | epsilon: 0.025
n = 268 | components = 3
Summary
Initial components: 3 | Components after truncation: 2
Summary table
parameter mean sd q0.025 q0.500 q0.975
weights[1] 0.783 0.016 0.759 0.774 0.799
weights[2] 0.17 0.012 0.156 0.167 0.201
alpha 1.004 0.566 0.253 1.022 2.199
beta_tail_scale[1] 0.039 0.061 -0.097 0.049 0.153
beta_tail_scale[2] -0.011 0.097 -0.165 -0.02 0.172
beta_tail_scale[3] 0.048 0.076 -0.09 0.041 0.193
beta_tail_scale[4] 1.061 0.12 0.837 1.06 1.24
tail_shape -0.02 0.019 -0.046 -0.028 0.01
threshold 7.173 0.105 6.827 7.204 7.204
shape1[1] 7.602 0.411 6.985 7.536 8.316
shape1[2] 5.569 1.256 2.787 5.513 7.623
shape2[1] 0.888 0.007 0.871 0.89 0.89
shape2[2] 0.27 0.12 0.112 0.289 0.452
beta_loc[1,1] 0 0 0 0 0
beta_loc[1,2] 0 0 0 0 0
beta_loc[1,3] 0 0 0 0 0
beta_loc[1,4] 0 0 0 0 0
beta_loc[2,1] 0 0 0 0 0
beta_loc[2,2] 0 0 0 0 0
beta_loc[2,3] 0 0 0 0 0
beta_loc[2,4] 0 0 0 0 0
beta_scale[1,1] -0.303 0.122 -0.518 -0.299 -0.095
beta_scale[1,2] -0.303 0.099 -0.468 -0.31 -0.09
beta_scale[1,3] 0.13 0.064 0.025 0.133 0.243
beta_scale[1,4] -0.13 0.133 -0.367 -0.14 0.089
beta_scale[2,1] 0.706 1.551 -1.923 0.563 3.96
beta_scale[2,2] 0.037 1.654 -3.141 0.081 3.161
beta_scale[2,3] -0.626 1.069 -2.831 -0.828 1.321
beta_scale[2,4] 1.32 2.616 -3.322 1.316 6.291
Code
Posterior mean parameters (causal)
[ps]
Posterior mean parameters
$beta
[1] 0.1103 0.4739 0.3935 -0.1501 0.1015
[treated]
Posterior mean parameters
$alpha
[1] 1.004
$w
[1] 0.7825 0.1704
$beta_loc
PropScore x1 x2 x3 x4
comp1 0 0 0 0 0
comp2 0 0 0 0 0
$beta_scale
PropScore x1 x2 x3 x4
comp1 0.730200 -0.3029 -0.30320 0.1301 -0.1296
comp2 0.002732 0.7057 0.03679 -0.6260 1.3200
$shape1
[1] 7.602 5.569
$shape2
[1] 0.8878 0.2705
$threshold
[1] 7.173
$beta_tail_scale
x1 x2 x3 x4
overall 0.03919 -0.01077 0.04789 1.061
$tail_shape
[1] -0.02018
[control]
Posterior mean parameters
$alpha
[1] 0.1923
$w
[1] 1
$beta_loc
PropScore x1 x2 x3 x4
comp1 2.473 -0.9426 0.1194 1.118 4.761
$beta_scale
PropScore x1 x2 x3 x4
comp1 0.08794 -0.1462 -0.278 -0.2206 -1.082
$shape1
[1] 5.2
$shape2
[1] 0.7054
$threshold
[1] 9.492
$beta_tail_scale
x1 x2 x3 x4
overall 0.08891 0.05928 0.023 0.3646
$tail_shape
[1] -0.2335
Code
plot (fit_sb_crp, family = plot_family_sb_crp_ex14)
=== treated ===
=== traceplot ===
=== control ===
=== traceplot ===
Code
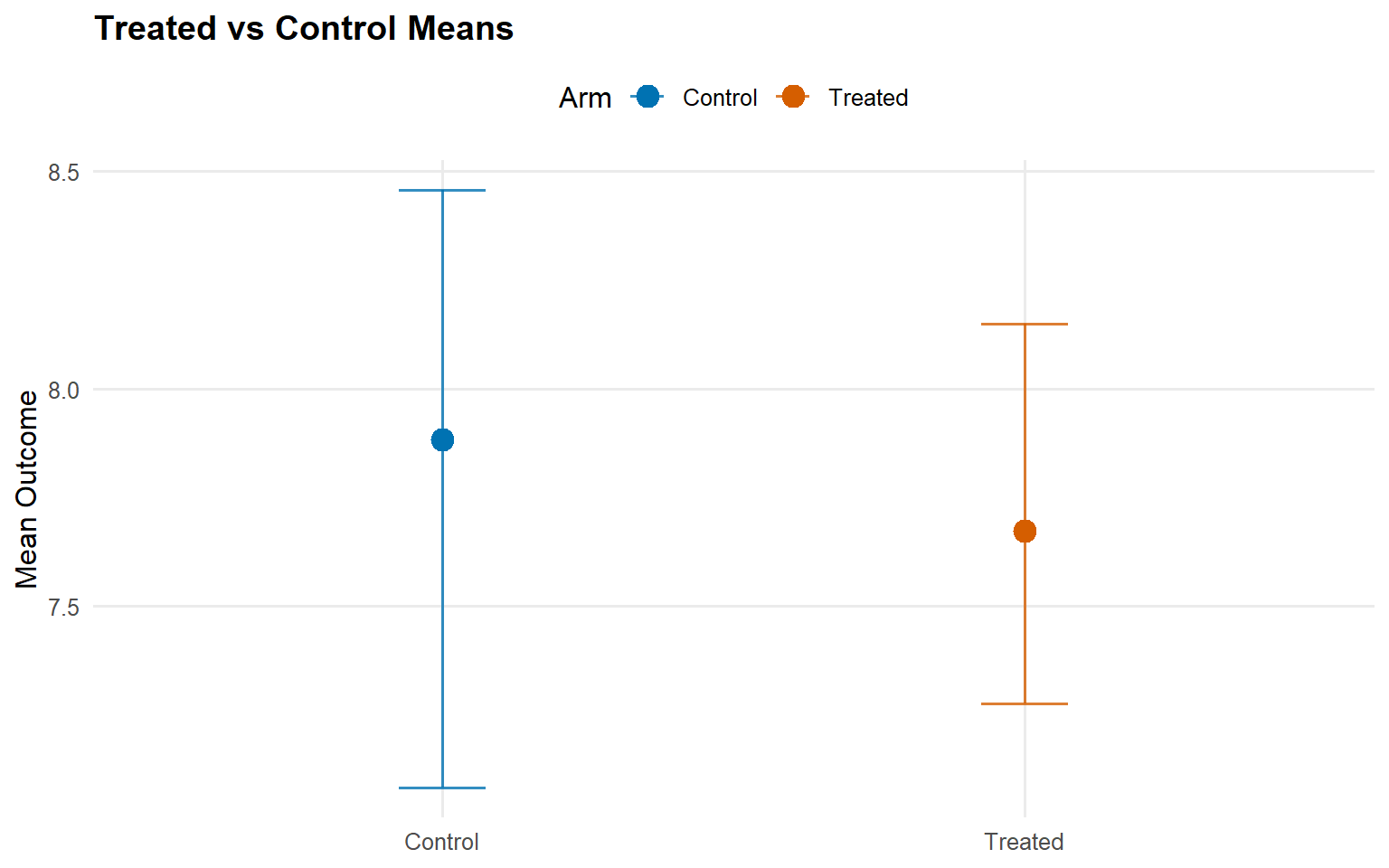
<- predict (fit_sb_crp, newdata = x_eval, type = "mean" ,interval = "credible" , nsim_mean = nsim_mean_ex14, workers = predict_workers_ex14)plot (pred_mean_sb_crp)
Code
<- predict (fit_sb_crp, newdata = x_eval, type = "quantile" ,p = 0.5 , interval = "credible" , workers = predict_workers_ex14)plot (pred_q_sb_crp)
Code
<- predict (fit_sb_crp, newdata = x_eval, y = y_eval,type = "density" , interval = "credible" , workers = predict_workers_ex14)plot (pred_d_sb_crp)
Code
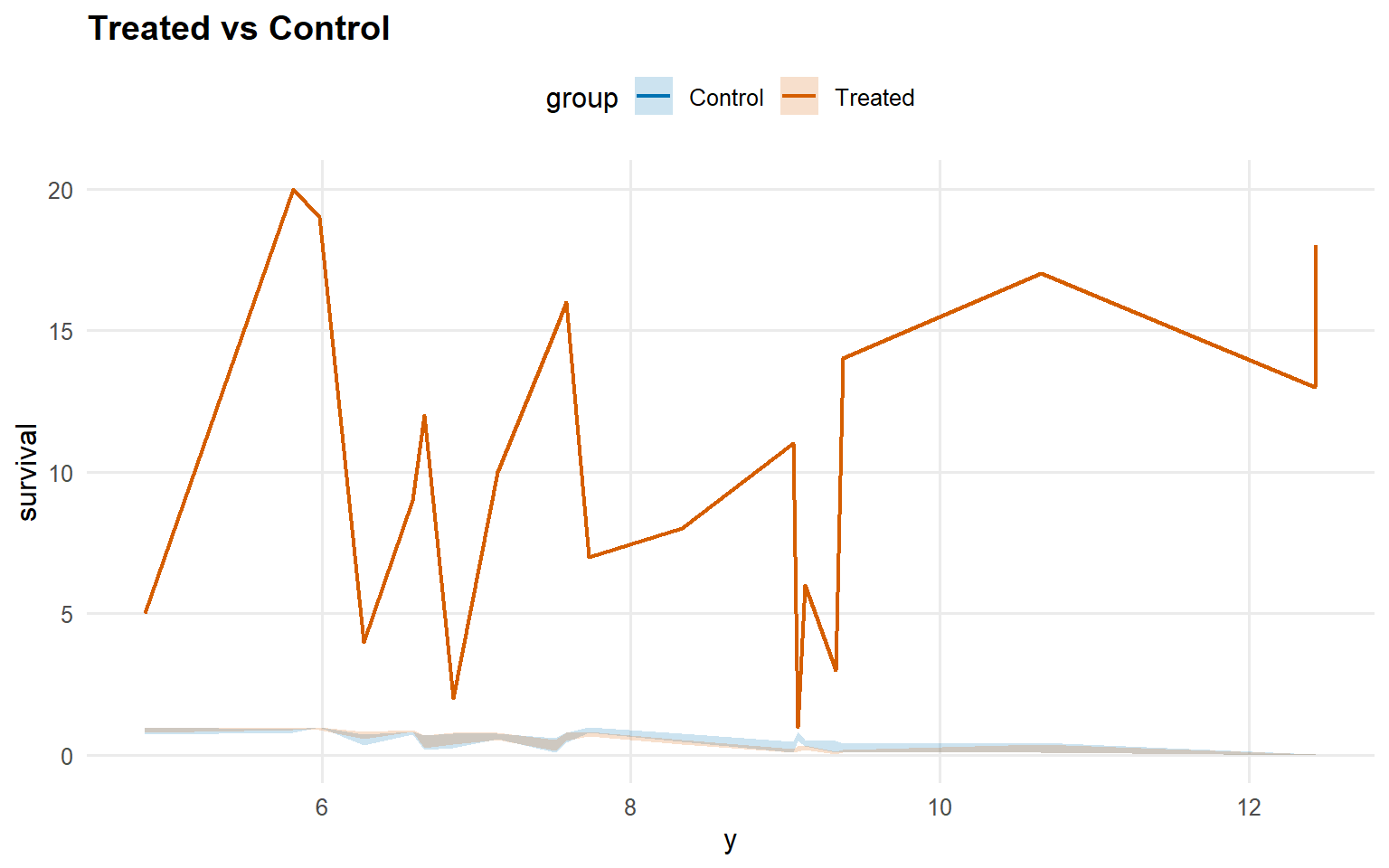
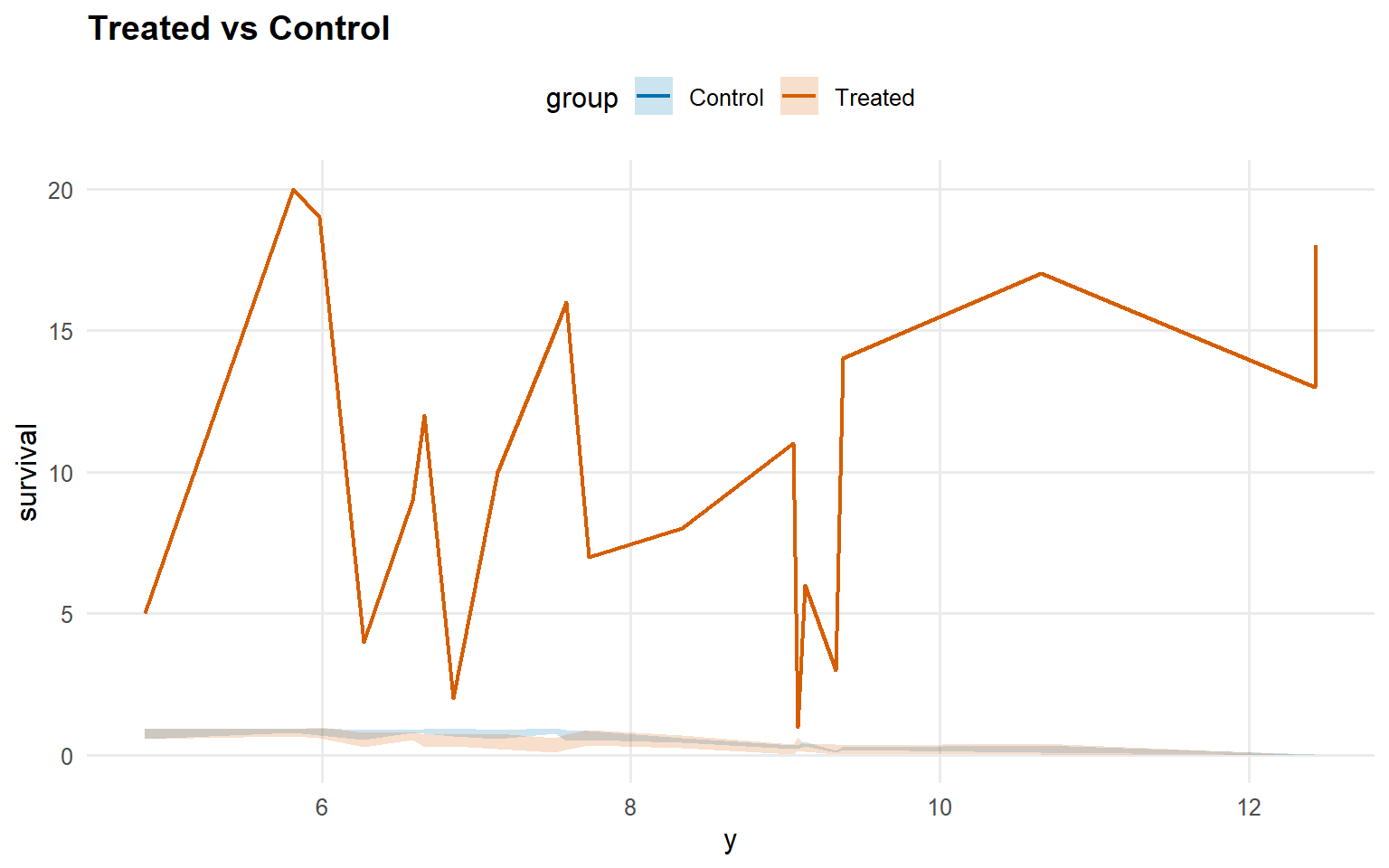
<- predict (fit_sb_crp, newdata = x_eval, y = y_eval,type = "survival" , interval = "credible" , workers = predict_workers_ex14)plot (pred_surv_sb_crp)
Code
<- ate (fit_sb_crp, interval = "credible" , nsim_mean = nsim_mean_ex14)plot (ate_sb_crp)
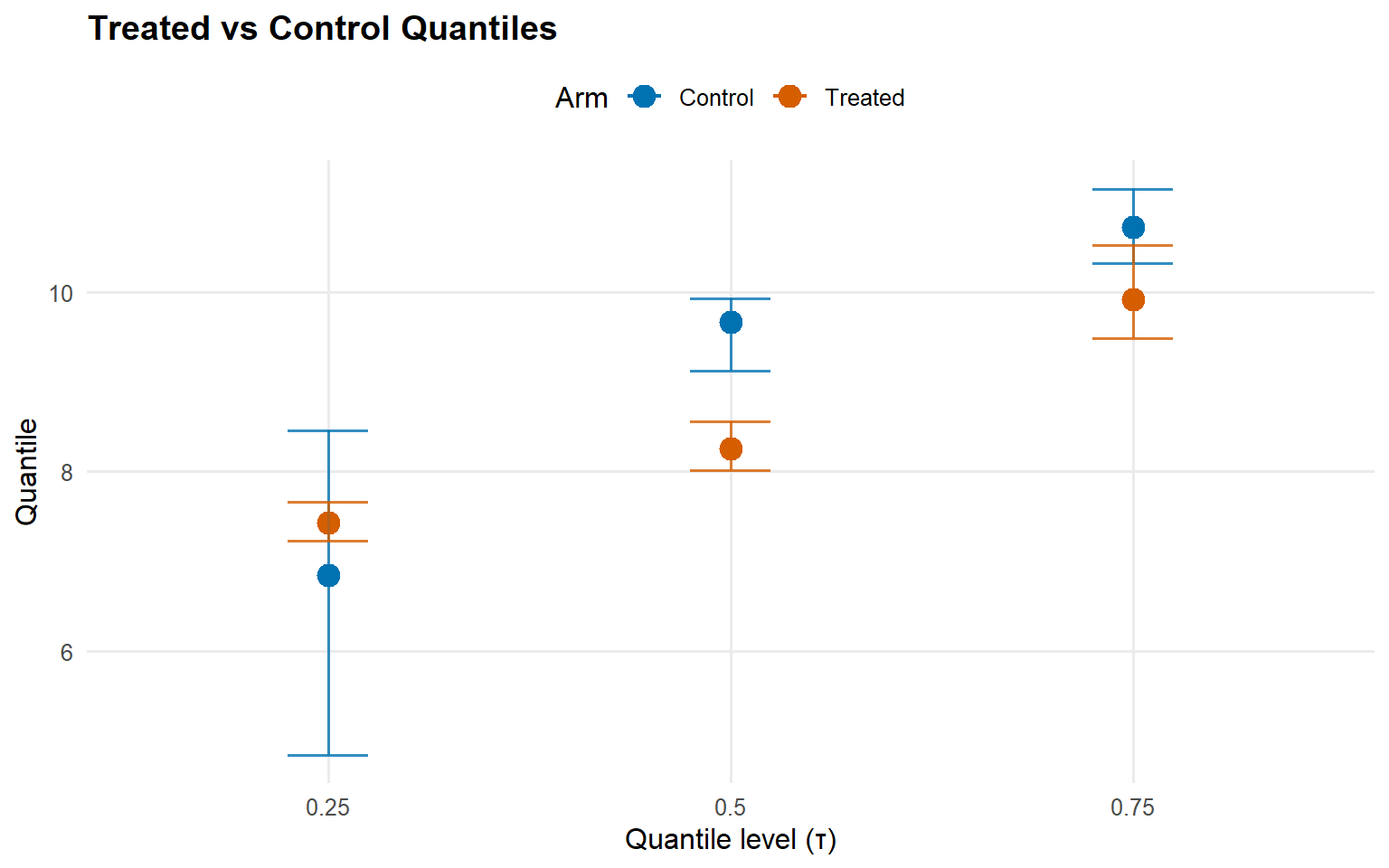
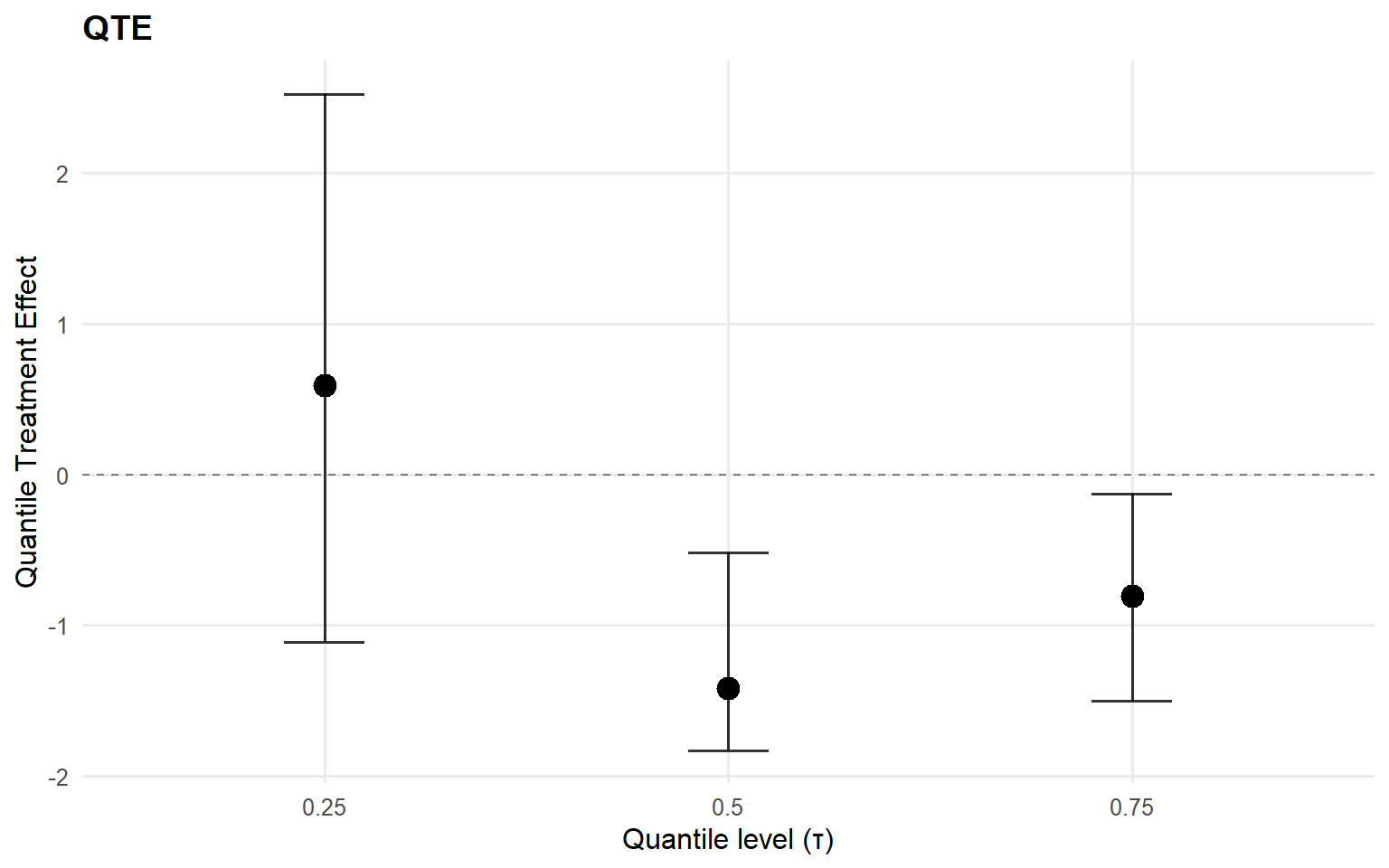
Code
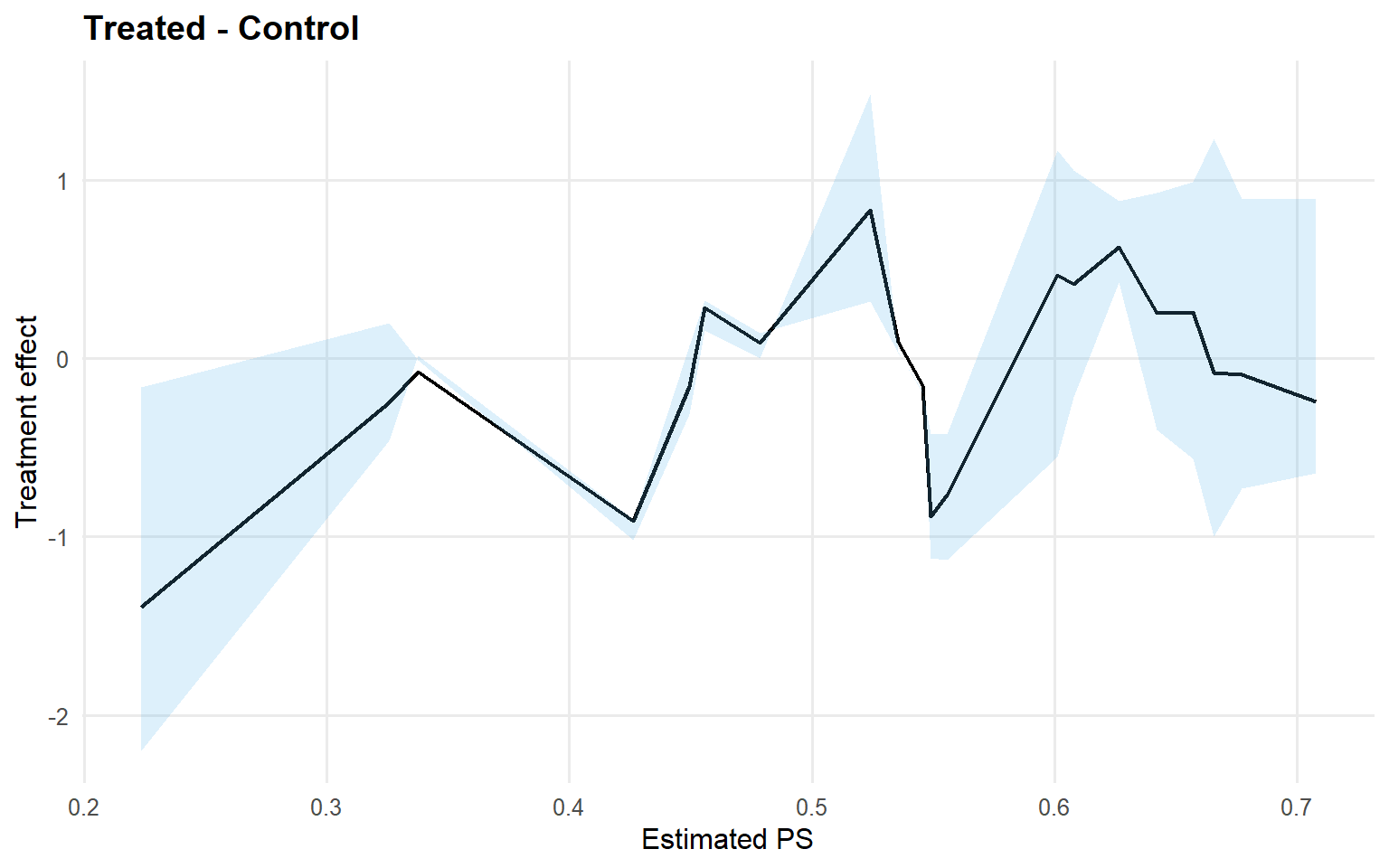
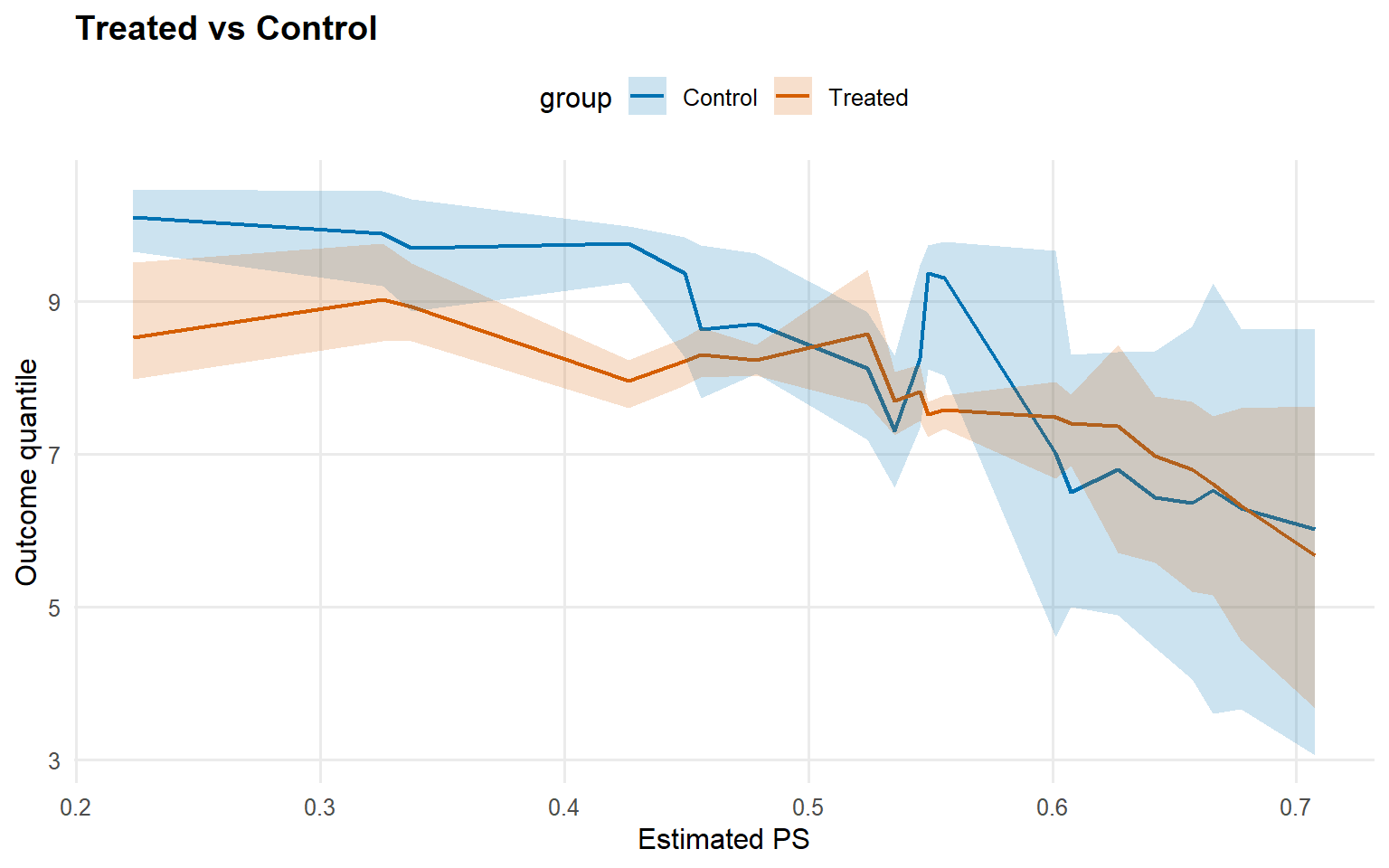
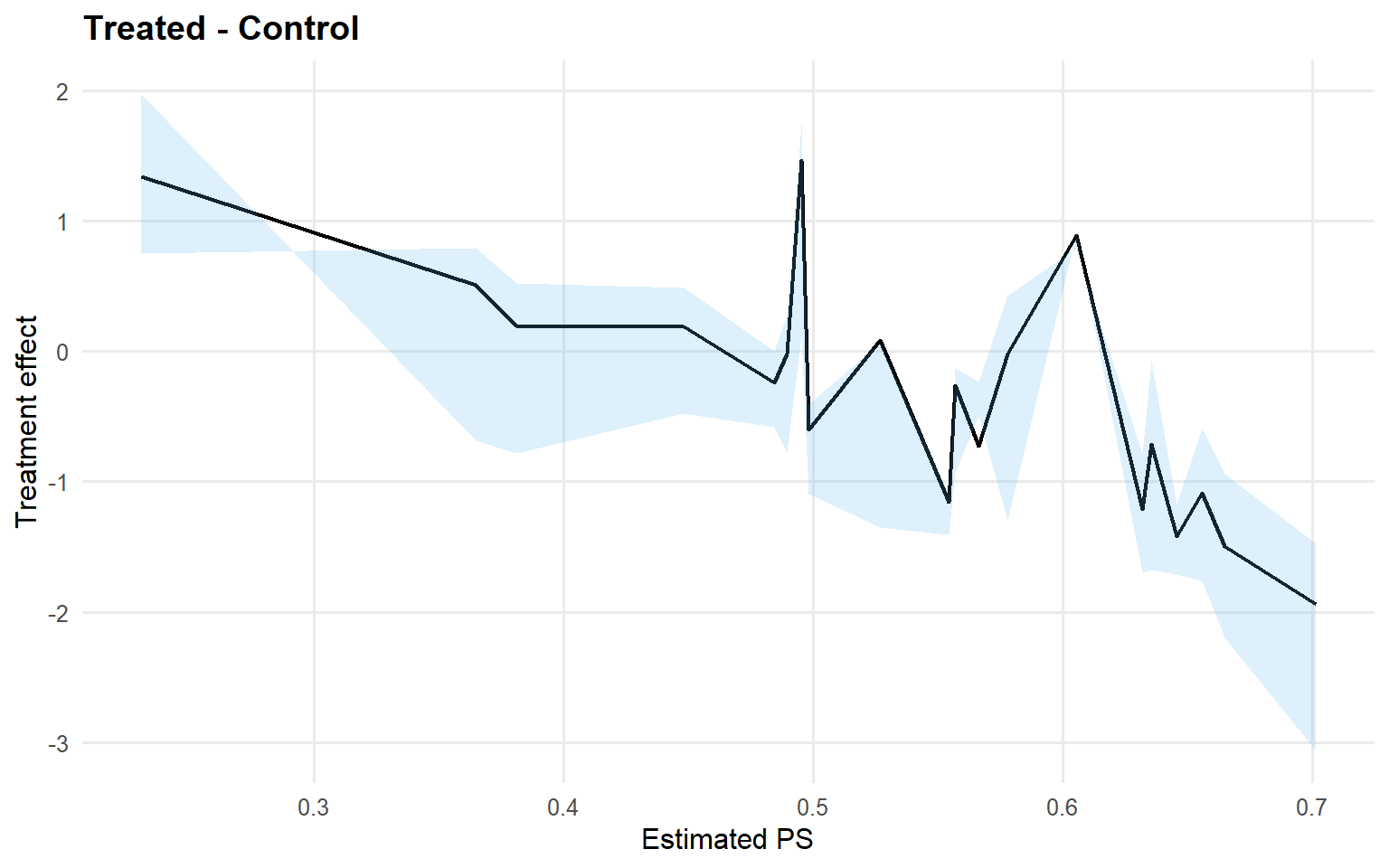
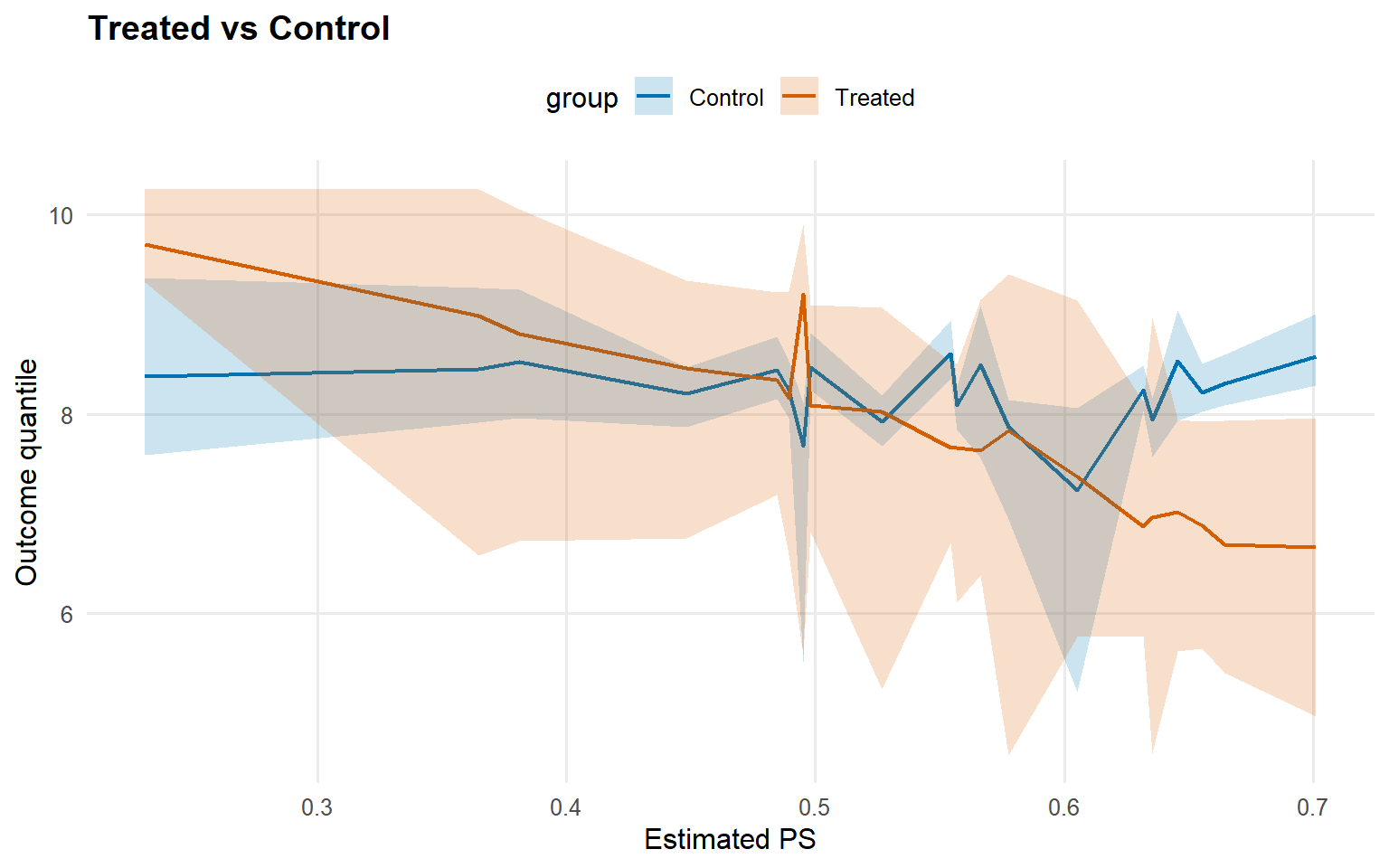
<- qte (fit_sb_crp, probs = c (0.25 , 0.5 , 0.75 ), interval = "credible" )plot (qte_sb_crp)
Model B: Treated CRP, Control SB (Amoroso + GPD)
Code
<- bundle (y = y,treat = A,X = X,kernel = c ("amoroso" , "amoroso" ),backend = c ("crp" , "sb" ),PS = "logit" ,GPD = c (TRUE , TRUE ),components = components_ex14,param_specs = param_specs_gpd,mcmc_outcome = gpd_mcmc_ex14_b,mcmc_ps = mcmc_ps_ex14_b
CausalMixGPD causal bundle
PS model: Bayesian logit (A | X)
Field Treated Control
Backend Chinese Restaurant Process Stick-Breaking Process
Kernel amoroso amoroso
Components 3 3
GPD tail TRUE TRUE
Epsilon 0.025 0.025
Outcome PS included: TRUE
n (control) = 232 | n (treated) = 268
Code
CausalMixGPD causal bundle summary
CausalMixGPD causal bundle
PS model: Bayesian logit (A | X)
Field Treated Control
Backend Chinese Restaurant Process Stick-Breaking Process
Kernel amoroso amoroso
Components 3 3
GPD tail TRUE TRUE
Epsilon 0.025 0.025
Outcome PS included: TRUE
n (control) = 232 | n (treated) = 268
Code
<- load_or_fit (quiet_mcmc (dpmgpd.causal (bundle_crp_sb, mcmc = causal_run_mcmc))summary (fit_crp_sb)
-- PS fit --
CausalMixGPD PS fit summary
model: logit
n = 500 | predictors = 5
Monitors: beta
Summary table
parameter mean sd q0.025 q0.500 q0.975
beta[1] 0.021 0.082 -0.056 -0.005 0.144
beta[2] 0.458 0.082 0.324 0.451 0.607
beta[3] 0.27 0.197 -0.042 0.216 0.544
beta[4] -0.149 0.065 -0.303 -0.107 -0.07
beta[5] 0.277 0.149 -0.228 0.348 0.48
-- Outcome fits --
[control]
MixGPD summary | backend: Stick-Breaking Process | kernel: Amoroso Distribution | GPD tail: TRUE | epsilon: 0.025
n = 232 | components = 3
Summary
Initial components: 3 | Components after truncation: 3
Summary table
parameter mean sd q0.025 q0.500 q0.975
weights[1] 0.493 0.014 0.487 0.487 0.527
weights[2] 0.452 0.03 0.409 0.474 0.474
weights[3] 0.055 0.023 0.038 0.038 0.099
alpha 1.831 0.884 0.563 1.628 3.332
beta_tail_scale[1] -0.053 0.056 -0.16 -0.052 0.052
beta_tail_scale[2] -0.028 0.101 -0.185 -0.028 0.14
beta_tail_scale[3] -0.045 0.049 -0.133 -0.04 0.033
beta_tail_scale[4] 0.889 0.142 0.65 0.905 1.166
tail_shape -0.077 0.022 -0.134 -0.084 -0.059
threshold 7.544 0 7.544 7.544 7.544
shape1[1] 5.107 0.924 4.032 4.943 7.146
shape1[2] 7.159 1.559 5.078 7.293 9.654
shape1[3] 2.642 1.475 0.238 2.694 5.982
shape2[1] 0.47 0.071 0.382 0.508 0.536
shape2[2] 0.985 0.028 0.955 0.971 1.016
shape2[3] 1.119 1.165 0.131 0.495 3.515
beta_loc[1,1] 0 0 0 0 0
beta_loc[1,2] 0 0 0 0 0
beta_loc[1,3] 0 0 0 0 0
beta_loc[1,4] 0 0 0 0 0
beta_loc[2,1] 0 0 0 0 0
beta_loc[2,2] 0 0 0 0 0
beta_loc[2,3] 0 0 0 0 0
beta_loc[2,4] 0 0 0 0 0
beta_loc[3,1] 0 0 0 0 0
beta_loc[3,2] 0 0 0 0 0
beta_loc[3,3] 0 0 0 0 0
beta_loc[3,4] 0 0 0 0 0
beta_scale[1,1] -0.263 0.47 -1.084 -0.277 0.76
beta_scale[1,2] 0.429 0.735 -0.884 0.354 1.616
beta_scale[1,3] -0.139 0.438 -0.795 -0.267 0.72
beta_scale[1,4] 2.034 1.116 0.011 1.949 4.311
beta_scale[2,1] 0.083 0.106 -0.15 0.072 0.259
beta_scale[2,2] 0.051 0.107 -0.125 0.041 0.26
beta_scale[2,3] -0.065 0.063 -0.194 -0.063 0.049
beta_scale[2,4] 0.365 0.372 -0.275 0.254 0.993
beta_scale[3,1] -0.287 2.184 -4.546 -0.539 4.407
beta_scale[3,2] -0.271 2.018 -3.948 -0.152 3.665
beta_scale[3,3] -0.127 1.554 -2.804 -0.451 3.516
beta_scale[3,4] 1.636 2.272 -2.587 1.848 5.503
[treated]
MixGPD summary | backend: Chinese Restaurant Process | kernel: Amoroso Distribution | GPD tail: TRUE | epsilon: 0.025
n = 268 | components = 3
Summary
Initial components: 3 | Components after truncation: 1
WAIC: 100140.928
lppd: -501.379 | pWAIC: 49569.085
Summary table
parameter mean sd q0.025 q0.500 q0.975
weights[1] 0.841 0.17 0.567 0.918 1
alpha 0.467 0.31 0.066 0.396 1.281
beta_tail_scale[1] 0.043 0.079 -0.094 0.047 0.19
beta_tail_scale[2] -0.153 0.121 -0.389 -0.144 0.088
beta_tail_scale[3] 0.201 0.104 -0.02 0.204 0.398
beta_tail_scale[4] 0.339 0.164 0.056 0.345 0.593
tail_shape -0.104 0.026 -0.173 -0.102 -0.076
threshold 9.028 0.072 9.003 9.003 9.292
shape1[1] 6.772 0.785 4.73 7.022 7.499
shape2[1] 0.875 0.018 0.861 0.861 0.908
beta_loc[1,1] 0.566 1.416 -2.103 1.346 1.968
beta_loc[1,2] -1.465 0.437 -2.125 -1.309 -0.617
beta_loc[1,3] -0.953 0.237 -1.258 -0.838 -0.61
beta_loc[1,4] 2.465 0.685 1.19 2.625 3.389
beta_scale[1,1] -0.298 0.223 -0.61 -0.359 0.21
beta_scale[1,2] 0.15 0.159 -0.151 0.186 0.448
beta_scale[1,3] 0.216 0.052 0.133 0.213 0.327
beta_scale[1,4] -0.528 0.234 -0.866 -0.558 0.05
Code
Posterior mean parameters (causal)
[ps]
Posterior mean parameters
$beta
[1] 0.02086 0.45850 0.27050 -0.14870 0.27740
[treated]
Posterior mean parameters
$alpha
[1] 0.4674
$w
[1] 0.8415
$beta_loc
PropScore x1 x2 x3 x4
comp1 0.8991 0.566 -1.465 -0.9532 2.465
$beta_scale
PropScore x1 x2 x3 x4
comp1 0.3048 -0.2978 0.1501 0.2158 -0.5283
$shape1
[1] 6.772
$shape2
[1] 0.8754
$threshold
[1] 9.028
$beta_tail_scale
x1 x2 x3 x4
overall 0.04303 -0.1532 0.2008 0.3389
$tail_shape
[1] -0.1039
[control]
Posterior mean parameters
$alpha
[1] 1.831
$w
[1] 0.49320 0.45200 0.05482
$beta_loc
PropScore x1 x2 x3 x4
comp1 0 0 0 0 0
comp2 0 0 0 0 0
comp3 0 0 0 0 0
$beta_scale
PropScore x1 x2 x3 x4
comp1 0.38570 -0.26260 0.4291 -0.13890 2.0340
comp2 -0.27440 0.08262 0.0505 -0.06538 0.3653
comp3 -0.02691 -0.28680 -0.2711 -0.12710 1.6360
$shape1
[1] 5.107 7.159 2.642
$shape2
[1] 0.4696 0.9845 1.1190
$threshold
[1] 7.544
$beta_tail_scale
x1 x2 x3 x4
overall -0.05256 -0.02801 -0.04545 0.8895
$tail_shape
[1] -0.07651
Code
plot (fit_crp_sb, family = plot_family_crp_sb_ex14)
=== treated ===
=== density ===
=== control ===
=== density ===
Code
<- predict (fit_crp_sb, newdata = x_eval, type = "mean" ,interval = "credible" , nsim_mean = nsim_mean_ex14, workers = predict_workers_ex14)plot (pred_mean_crp_sb)
Code
<- predict (fit_crp_sb, newdata = x_eval, type = "quantile" ,p = 0.5 , interval = "credible" , workers = predict_workers_ex14)plot (pred_q_crp_sb)
Code
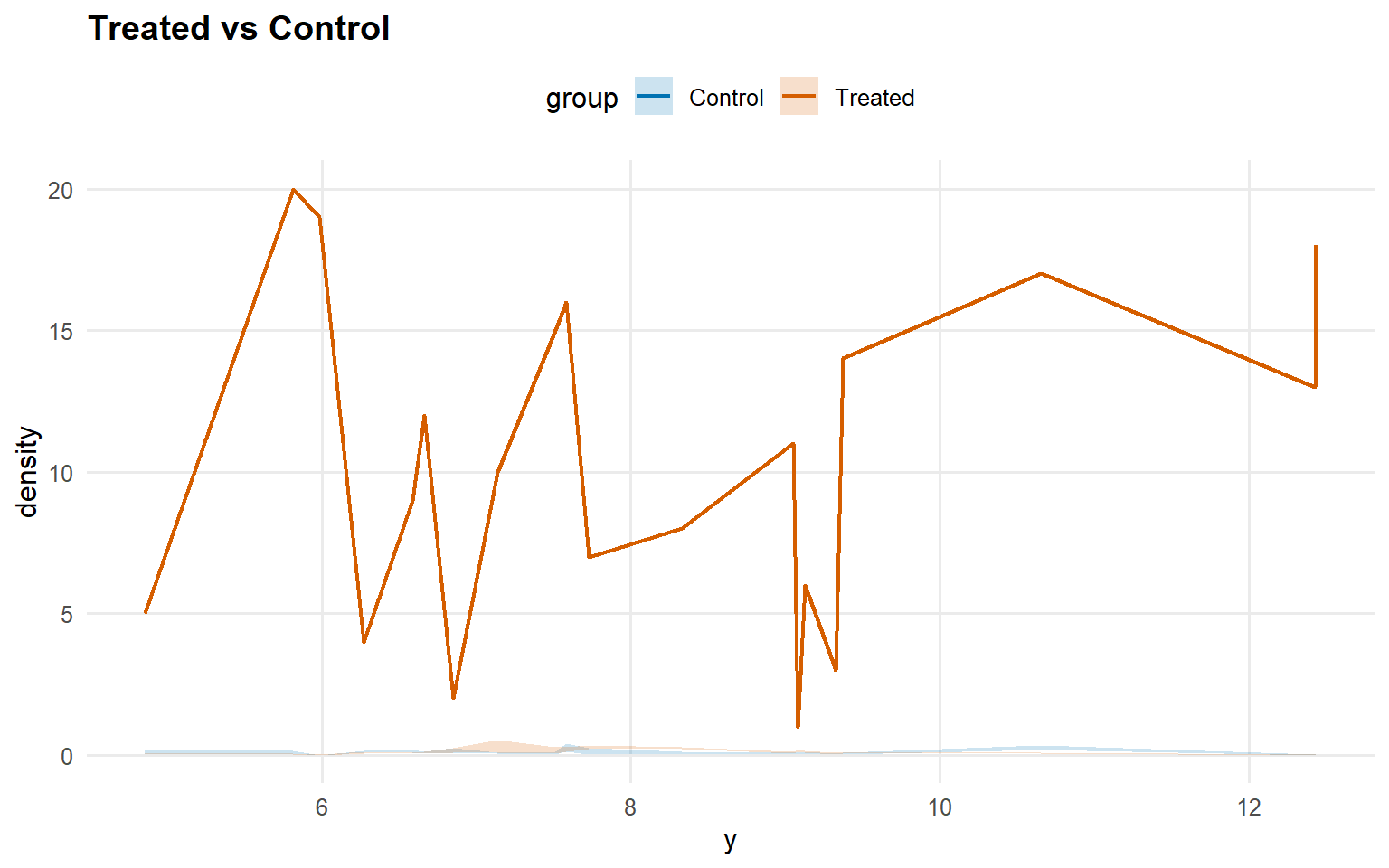
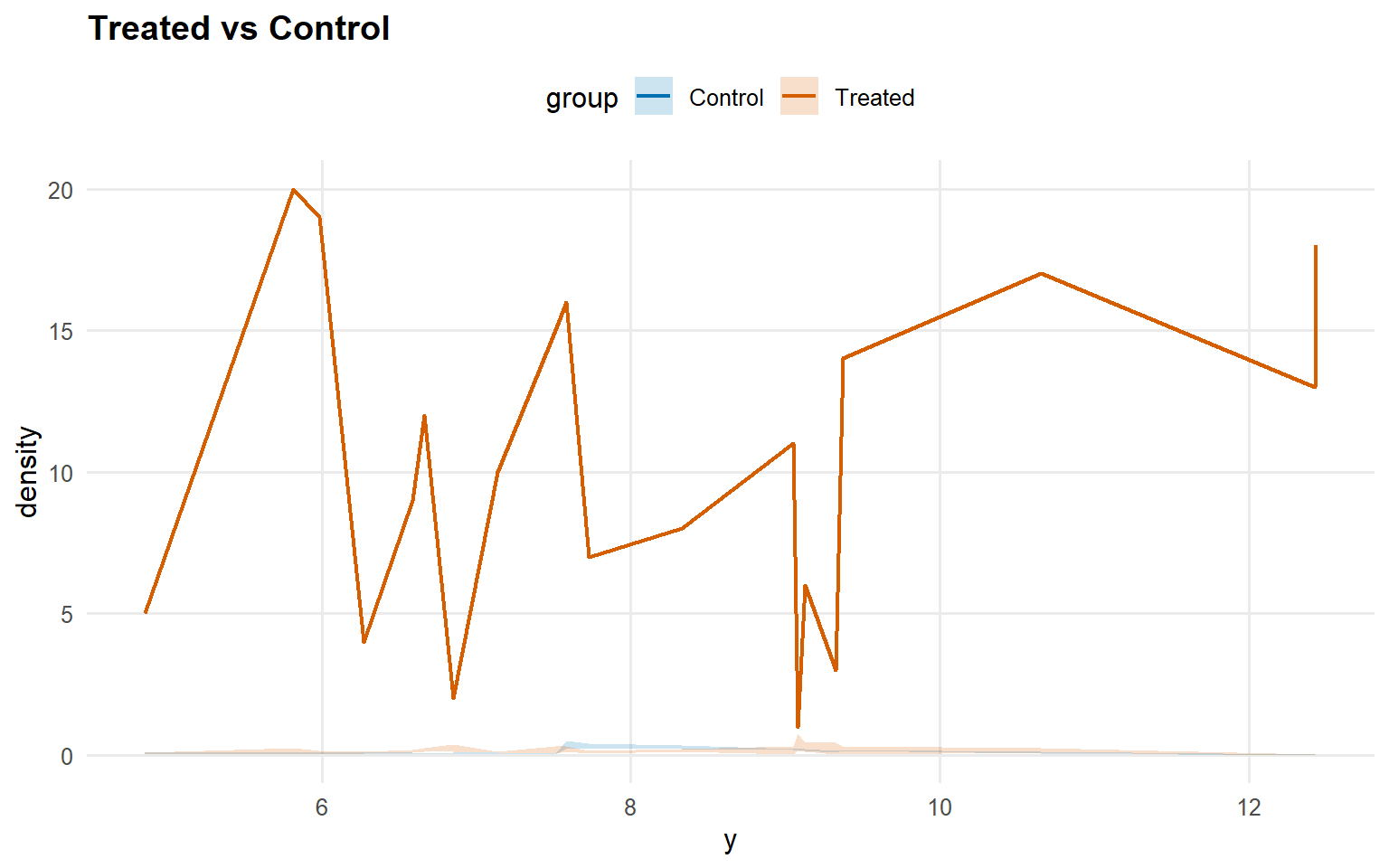
<- predict (fit_crp_sb, newdata = x_eval, y = y_eval,type = "density" , interval = "credible" , workers = predict_workers_ex14)plot (pred_d_crp_sb)
Code
<- predict (fit_crp_sb, newdata = x_eval, y = y_eval,type = "survival" , interval = "credible" , workers = predict_workers_ex14)plot (pred_surv_crp_sb)
Code
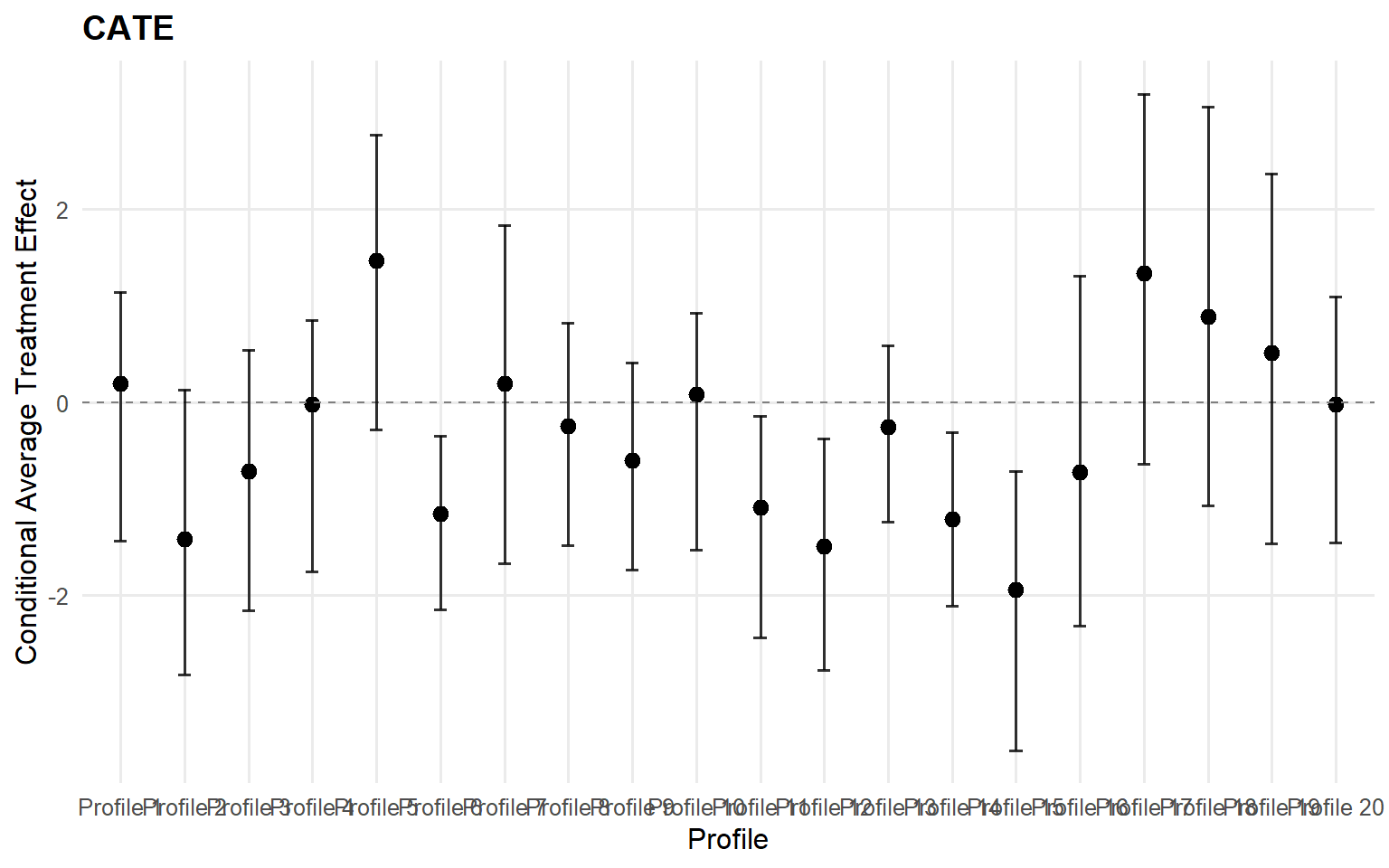
<- cate (fit_crp_sb, newdata = x_eval,interval = "credible" , nsim_mean = nsim_mean_ex14)head (format_cate_table (cate_crp_sb))
id index estimate lower upper
1 1 1 0.18953974 -1.4347426 1.1397729
2 2 1 -1.41838105 -2.8257883 0.1279135
3 3 1 -0.71391510 -2.1527255 0.5422384
4 4 1 -0.02181351 -1.7503993 0.8510921
5 5 1 1.46568595 -0.2840095 2.7674652
6 6 1 -1.15685357 -2.1452202 -0.3455291
Code
Code
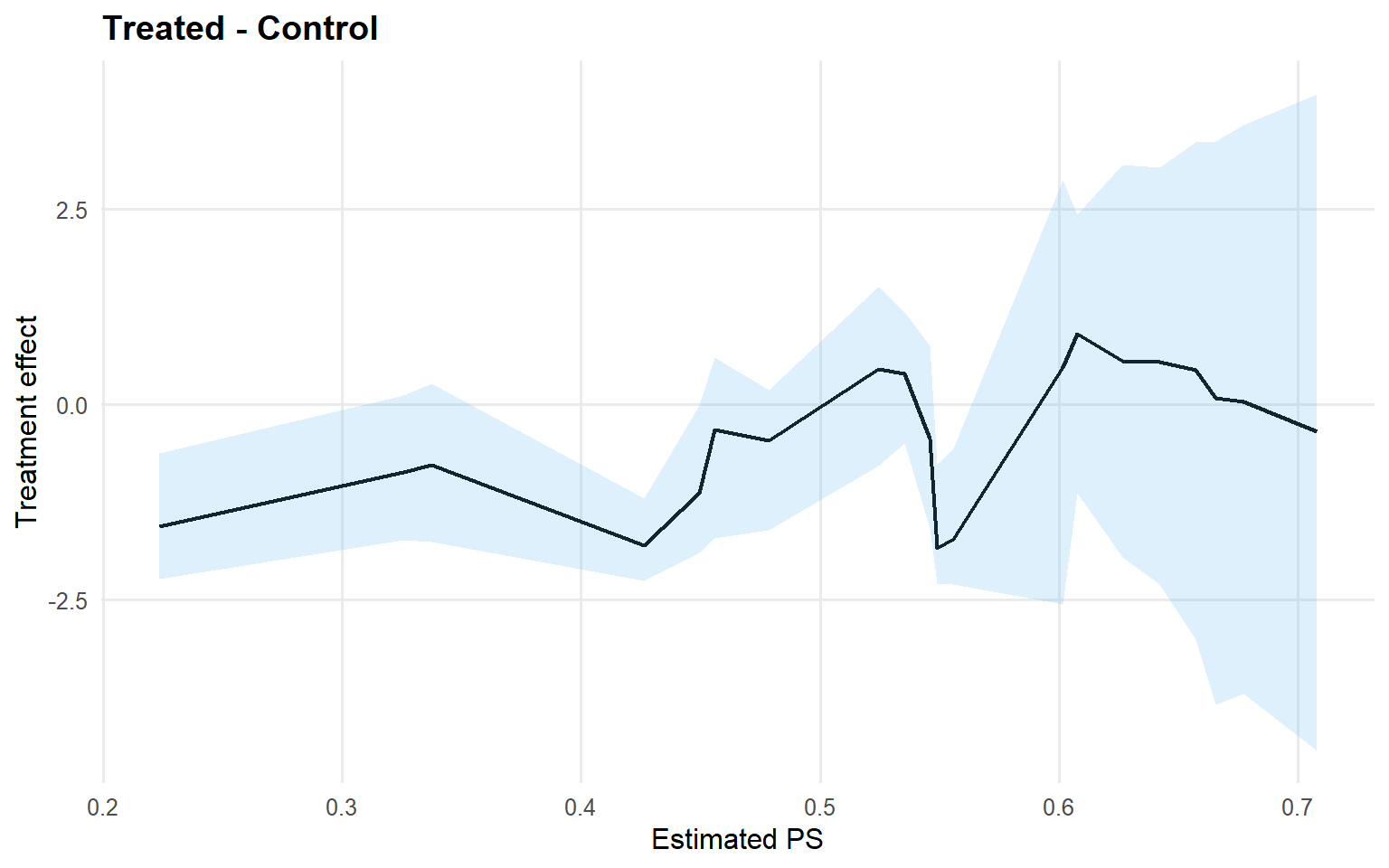
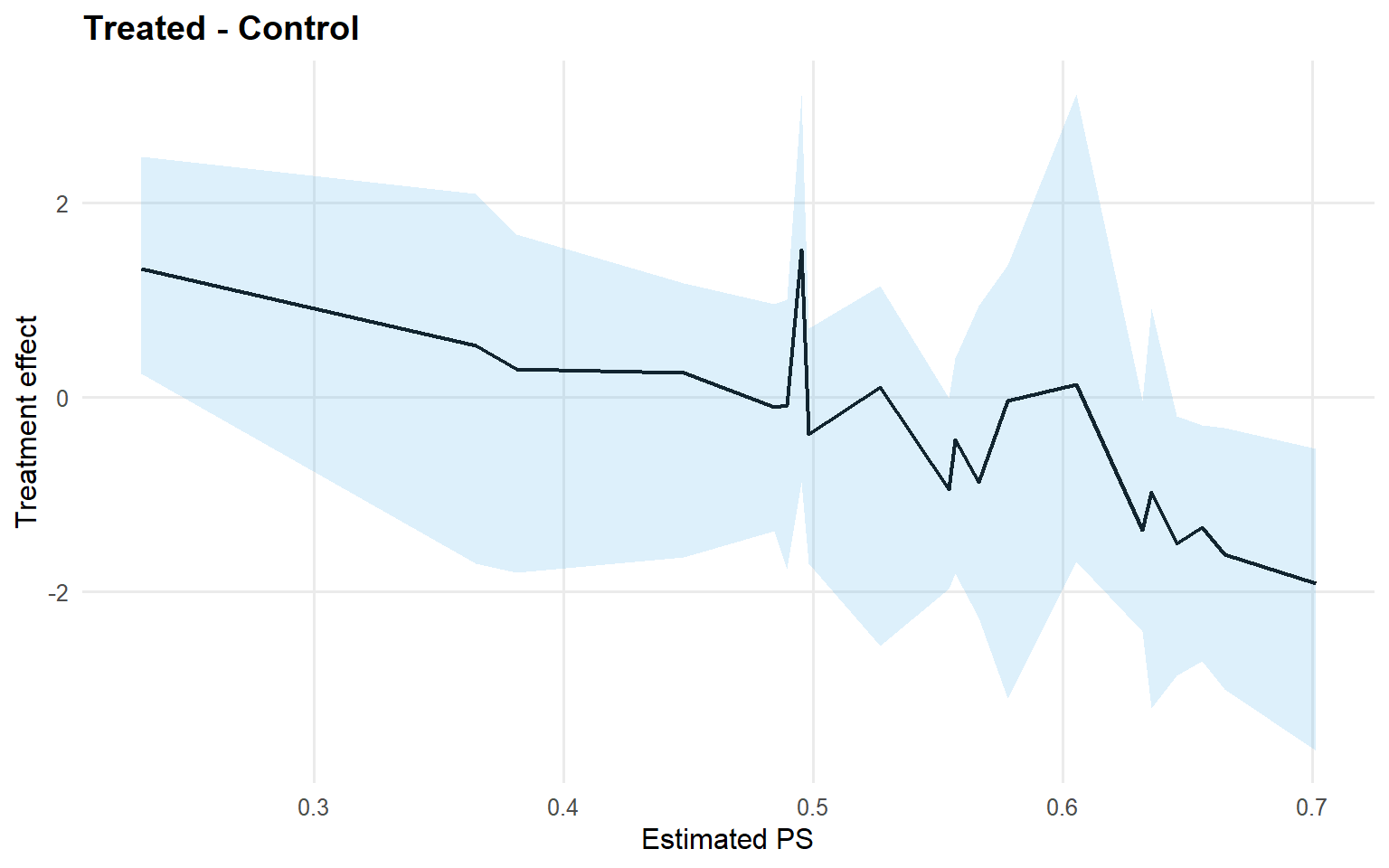
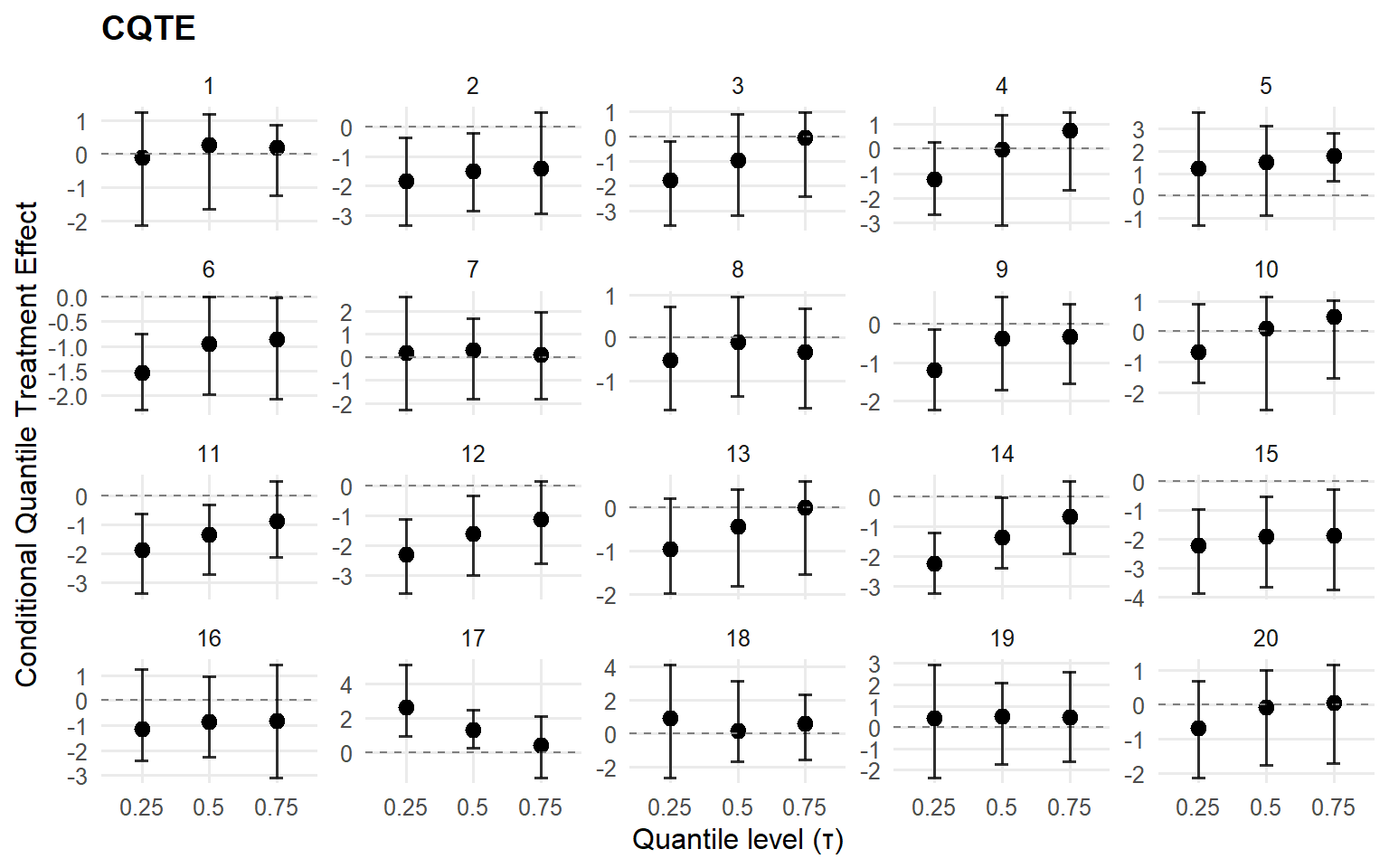
<- cqte (fit_crp_sb, probs = c (0.25 , 0.5 , 0.75 ),newdata = x_eval, interval = "credible" )head (format_cqte_table (cqte_crp_sb))
id index estimate lower upper
1 1 1 -0.1229123 -2.135624 1.2259354
2 1 2 0.2534521 -1.649174 1.1700091
3 1 3 0.1790422 -1.239670 0.8398795
4 2 1 -1.8316943 -3.343504 -0.3658125
5 2 2 -1.5103598 -2.865960 -0.1963388
6 2 3 -1.4044933 -2.934697 0.5036670
Code
Prereqs
Required packages and data for this page are listed in the setup chunks above.
Outputs
This page renders model fits, diagnostics, and summary artifacts generated by package APIs.
Interpretation
Canonical concept page: 03 Causal Inference Objects
Treat this page as an application/example view and use the canonical page for core definitions.
Next
Continue to the linked canonical concept page, then return for implementation-specific details.