Website workflow note. This page reflects the current exported API and recommended wrapper-first usage. Last updated: 2026-02-19 .
For the full package narrative, see the main package vignettes (basic, unconditional, conditional, and causal).
Conditional DPmix: Stick-Breaking Backend
Purpose : Replace the CRP backend with stick-breaking truncation while keeping the covariate-dependent bulk structure. This demonstrates how fixed components interplay with covariates.
What you’ll learn
How to run the conditional (y \mid X) workflow using the SB backend (fixed truncation).
How components controls effective mixture complexity in conditional fits.
How to compare kernels under the same SB setup using parallel bundles.
When to use this template
You want conditional density estimation with predictable runtime and a tunable complexity knob (components).
You want conditional modeling but do not need tail splicing (GPD = FALSE).
Next steps
Add tail augmentation for covariate-dependent extremes (ex08), or compare with CRP conditional (ex05).
Data Setup
Code
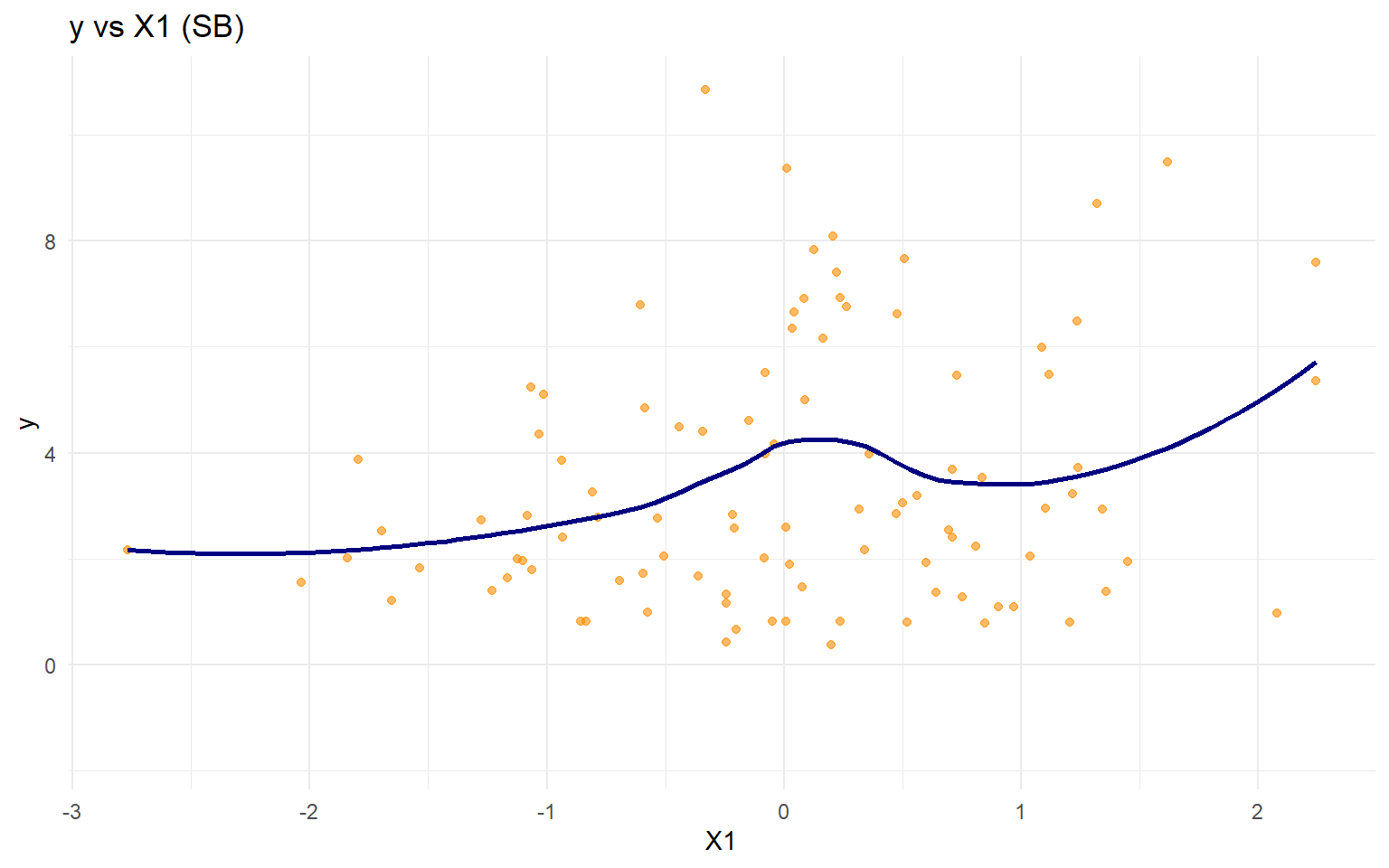
data ("nc_posX100_p3_k2" )<- nc_posX100_p3_k2$ y<- as.matrix (nc_posX100_p3_k2$ X)if (is.null (colnames (X))) {colnames (X) <- paste0 ("x" , seq_len (ncol (X)))<- tibble (statistic = c ("N" , "Mean" , "SD" , "Min" , "Max" ),value = c (length (y), mean (y), sd (y), min (y), max (y))ggplot (data.frame (y = y, x1 = X[, 1 ]), aes (x = x1, y = y)) + geom_point (alpha = 0.6 , color = "darkorange" ) + geom_smooth (method = "loess" , color = "navy" , fill = NA ) + labs (title = "y vs X1 (SB)" , x = "X1" , y = "y" ) + theme_minimal ()
# A tibble: 5 × 2
statistic value
<chr> <dbl>
1 N 100
2 Mean 3.45
3 SD 2.41
4 Min 0.377
5 Max 10.9
Model Specification
Code
<- bundle (y = y,X = X,kernel = "normal" ,backend = "sb" ,GPD = FALSE ,components = 5 ,mcmc = mcmc<- bundle (y = y,X = X,kernel = "cauchy" ,backend = "sb" ,GPD = FALSE ,components = 5 ,mcmc = mcmc
Running MCMC
Code
<- load_or_fit ("ex06-conditional-dpm-sb-fit_sb_normal" , dpmix (bundle_sb_normal))<- load_or_fit ("ex06-conditional-dpm-sb-fit_sb_cauchy" , dpmix (bundle_sb_cauchy))summary (fit_sb_normal)
MixGPD summary | backend: Stick-Breaking Process | kernel: Normal Distribution | GPD tail: FALSE | epsilon: 0.025
n = 100 | components = 5
Summary
Initial components: 5 | Components after truncation: 2
Summary table
parameter mean sd q0.025 q0.500 q0.975 ess
weights[1] 0.71 0.125 0.432 0.717 0.891 2.821
weights[2] 0.188 0.091 0.075 0.139 0.353 3.134
alpha 0.973 0.467 0.266 0.836 2.086 30.527
beta_mean[1, 1] -0.697 0.689 -2.269 -0.399 0.193 3.626
beta_mean[2, 1] 0.874 0.527 -0.155 1.016 1.891 3.601
beta_mean[3, 1] -0.21 0.482 -1.133 -0.228 0.599 11.882
beta_mean[4, 1] -0.86 0.553 -1.798 -0.897 0.353 6.33
beta_mean[5, 1] 0.552 0.522 -1.185 0.581 1.235 8.768
beta_mean[1, 2] 1.274 0.846 0.465 0.976 3.069 2.147
beta_mean[2, 2] 0.3 0.45 -0.736 0.399 1.115 8.059
beta_mean[3, 2] -0.982 0.425 -1.82 -0.838 -0.251 12.352
beta_mean[4, 2] -0.33 0.314 -1.136 -0.367 0.335 29.796
beta_mean[5, 2] 2.136 0.702 1.518 1.919 3.718 2.955
beta_mean[1, 3] -0.464 0.897 -2.007 -0.273 0.447 2.248
beta_mean[2, 3] 0.336 0.852 -0.594 0.263 1.843 1.602
beta_mean[3, 3] -1.703 0.564 -2.487 -1.858 -0.072 6.078
beta_mean[4, 3] 0.792 0.817 -0.046 0.432 2.963 2.573
beta_mean[5, 3] 1.737 0.812 0.548 1.701 3.785 3.443
sd[1] 0.053 0.01 0.036 0.053 0.073 52.239
sd[2] 0.794 1.107 0.037 0.234 3.846 22.413
Code
MixGPD summary | backend: Stick-Breaking Process | kernel: Cauchy Distribution | GPD tail: FALSE | epsilon: 0.025
n = 100 | components = 5
Summary
Initial components: 5 | Components after truncation: 3
Summary table
parameter mean sd q0.025 q0.500 q0.975 ess
weights[1] 0.435 0.078 0.297 0.447 0.573 8.109
weights[2] 0.285 0.049 0.189 0.3 0.35 10.355
weights[3] 0.17 0.046 0.083 0.172 0.258 11.74
alpha 1.381 0.846 0.319 1.133 3.513 17.696
beta_location[1, 1] 1.031 1.408 -1.795 1.507 2.522 1.503
beta_location[2, 1] 1.135 0.786 -0.182 1.183 2.12 4.936
beta_location[3, 1] 0.758 1.099 -1.514 0.772 2.525 3.277
beta_location[4, 1] 0.153 0.777 -0.631 -0.184 1.832 2.518
beta_location[5, 1] 0.091 0.661 -0.979 0.309 1.353 9.197
beta_location[1, 2] 0.532 0.927 -0.891 0.133 2.046 2.91
beta_location[2, 2] -1.785 1.241 -3.511 -1.88 0.311 2.29
beta_location[3, 2] -3.126 1.533 -4.652 -3.934 0.302 1.866
beta_location[4, 2] -1.304 1.338 -3.017 -1.371 1.447 2.479
beta_location[5, 2] 1.507 0.419 0.527 1.623 2.03 9.906
beta_location[1, 3] 1.058 0.827 -1.123 0.94 2.358 4.52
beta_location[2, 3] 1.654 1.804 -2.067 2.43 3.752 2.449
beta_location[3, 3] 1.364 1.264 -1.047 1.404 2.961 2.195
beta_location[4, 3] -1.01 1.051 -2.325 -1.35 1.259 1.961
beta_location[5, 3] -1.34 1.028 -3.261 -1.114 0.3 2.243
scale[1] 2.046 0.56 1.16 1.964 3.233 103.998
scale[2] 2.19 0.82 0.978 2.027 4.075 78.988
scale[3] 1.99 0.966 0.612 1.788 4.133 82.816
Code
<- params (fit_sb_normal)
Posterior mean parameters
$alpha
[1] 0.9726
$w
[1] 0.7103 0.1881
$beta_mean
x1 x2 x3
comp1 -0.6969 1.2740 -0.4642
comp2 0.8744 0.3004 0.3364
comp3 -0.2103 -0.9816 -1.7030
comp4 -0.8597 -0.3299 0.7915
comp5 0.5520 2.1360 1.7370
$sd
[1] 0.05346 0.79370
Conditional Predictive Density
Code
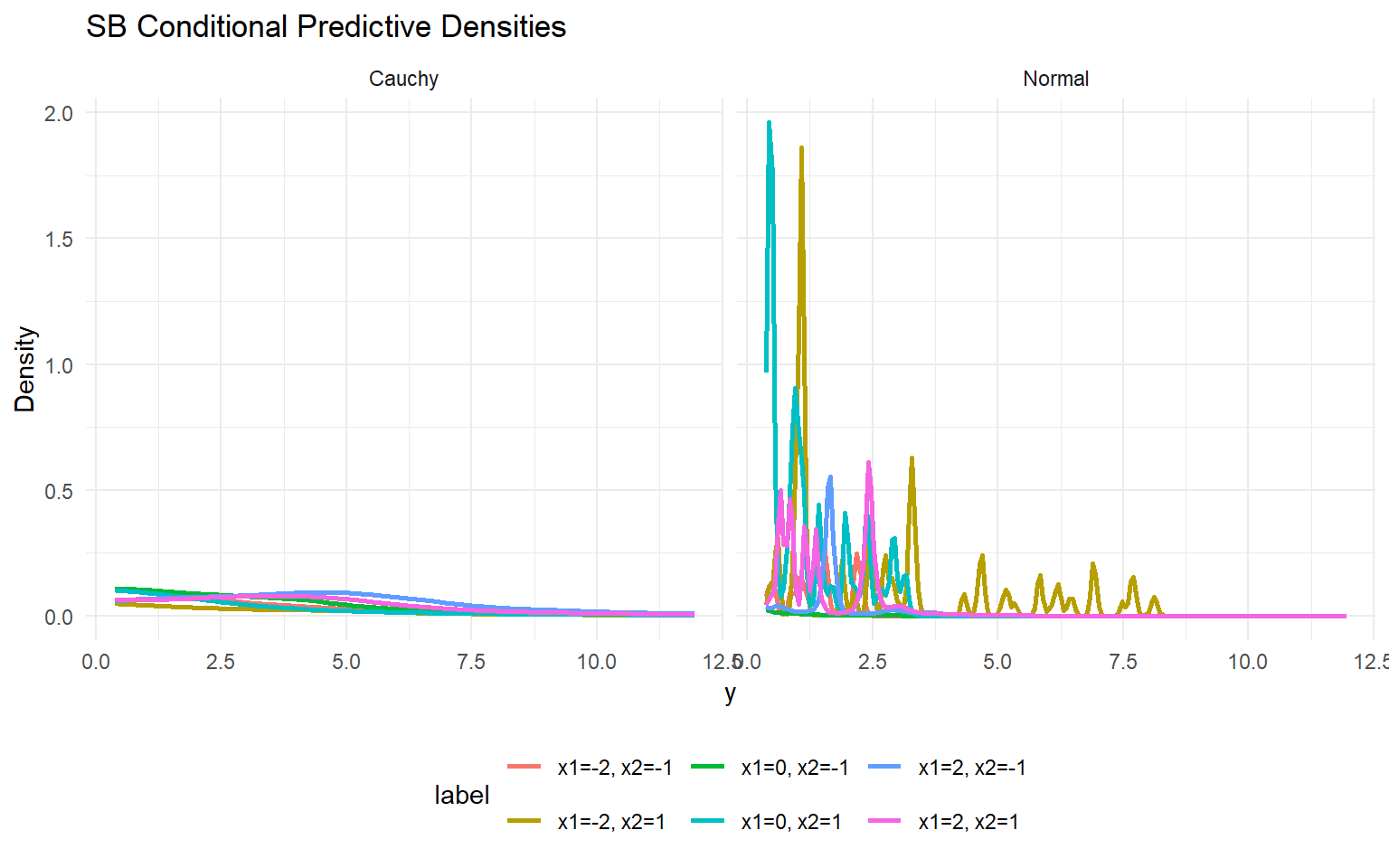
<- expand.grid (x1 = seq (- 2 , 2 , length.out = 3 ), x2 = c (- 1 , 1 ), x3 = 0 )colnames (X_new) <- colnames (X)<- max (min (y), .Machine$ double.eps)<- seq (y_min, max (y) * 1.1 , length.out = 200 )<- lapply (seq_len (nrow (X_new)), function (i) {<- predict (fit_sb_normal, newdata = as.matrix (X_new[i, , drop = FALSE ]), y = y_grid, type = "density" )data.frame (y = pred$ fit$ y,density = pred$ fit$ density,label = paste0 ("x1=" , round (X_new[i, "x1" ], 1 ), ", x2=" , X_new[i, "x2" ]),model = "Normal" <- lapply (seq_len (nrow (X_new)), function (i) {<- predict (fit_sb_cauchy, newdata = as.matrix (X_new[i, , drop = FALSE ]), y = y_grid, type = "density" )data.frame (y = pred$ fit$ y,density = pred$ fit$ density,label = paste0 ("x1=" , round (X_new[i, "x1" ], 1 ), ", x2=" , X_new[i, "x2" ]),model = "Cauchy" <- bind_rows (densities_normal, densities_cauchy)ggplot (df_cond, aes (x = y, y = density, color = label)) + geom_line (linewidth = 1 ) + facet_wrap (~ model) + labs (title = "SB Conditional Predictive Densities" , x = "y" , y = "Density" ) + theme_minimal () + theme (legend.position = "bottom" )
Quantile Drift with Covariates
Code
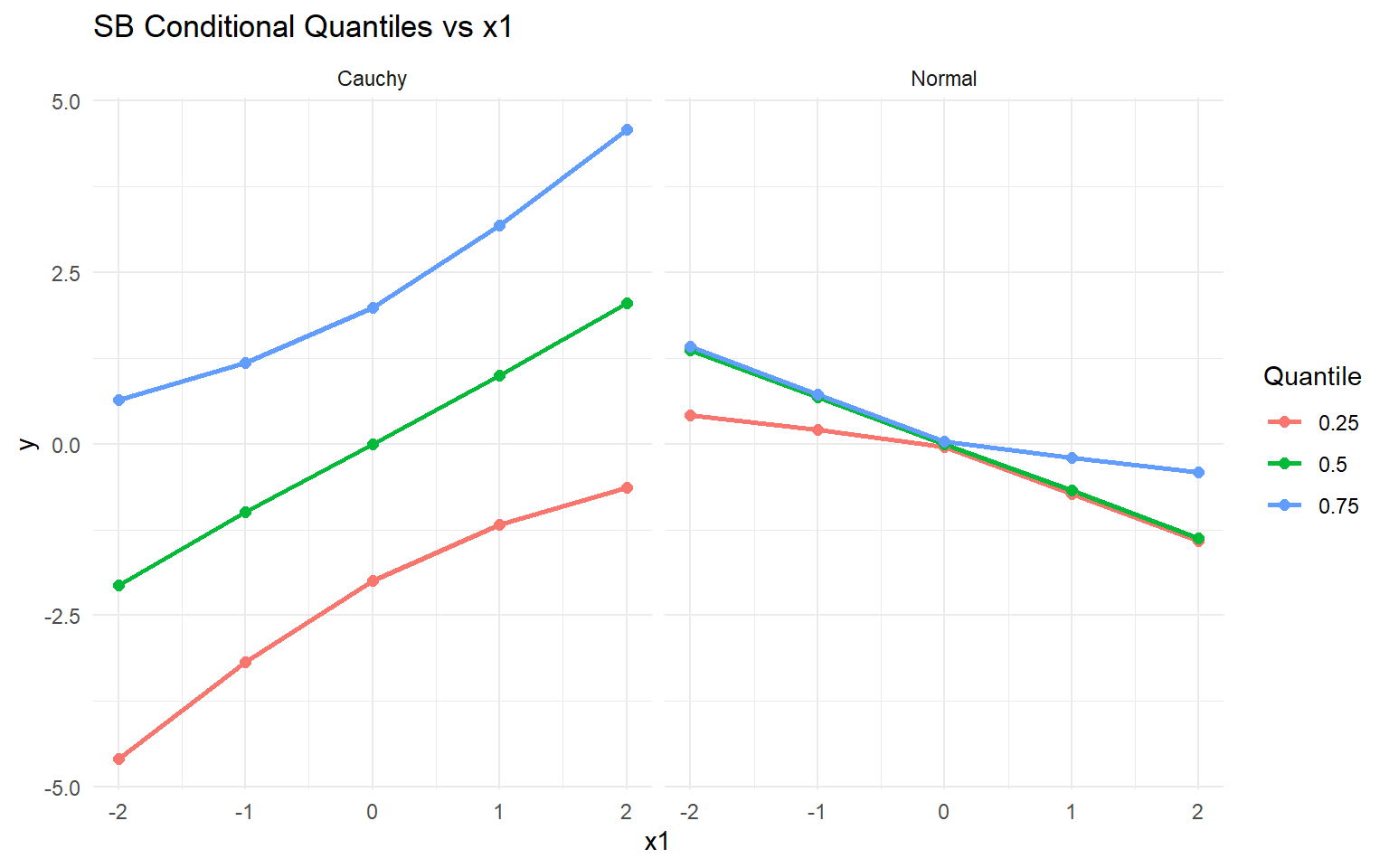
<- cbind (x1 = seq (- 2 , 2 , length.out = 5 ), x2 = 0 , x3 = 0 )colnames (X_eval) <- colnames (X)<- c (0.25 , 0.5 , 0.75 )<- predict (fit_sb_normal, newdata = as.matrix (X_eval), type = "quantile" , p = quant_probs)<- predict (fit_sb_cauchy, newdata = as.matrix (X_eval), type = "quantile" , p = quant_probs)<- pred_q_normal$ fit$ x1 <- X_eval[quant_df_normal$ id, "x1" ]$ model <- "Normal" <- pred_q_cauchy$ fit$ x1 <- X_eval[quant_df_cauchy$ id, "x1" ]$ model <- "Cauchy" bind_rows (quant_df_normal, quant_df_cauchy) %>% ggplot (aes (x = x1, y = estimate, color = factor (index), group = index)) + geom_line (linewidth = 1 ) + geom_point (size = 2 ) + facet_wrap (~ model) + labs (title = "SB Conditional Quantiles vs x1" , x = "x1" , y = "y" , color = "Quantile" ) + theme_minimal ()
Residuals & Diagnostics
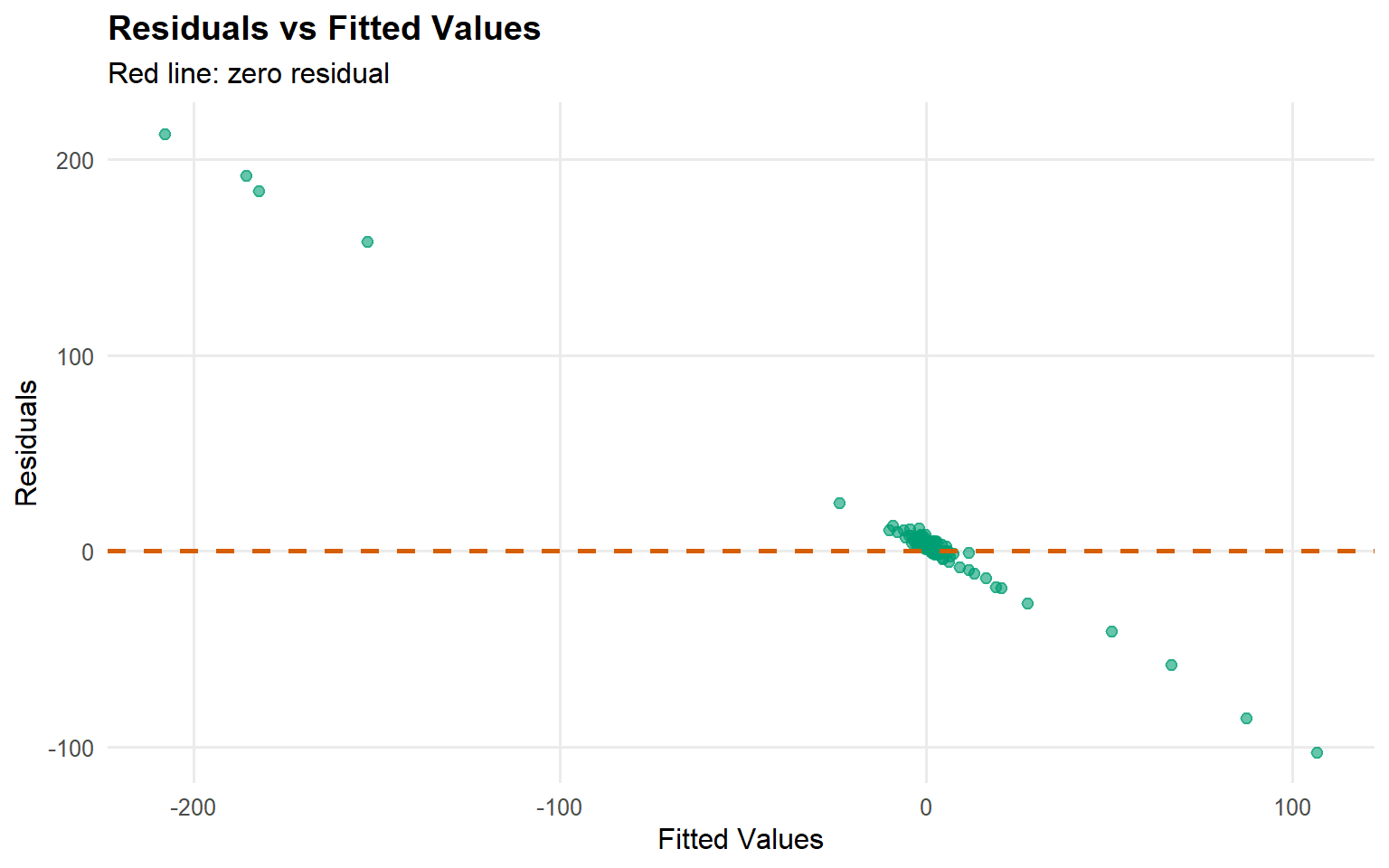
Code
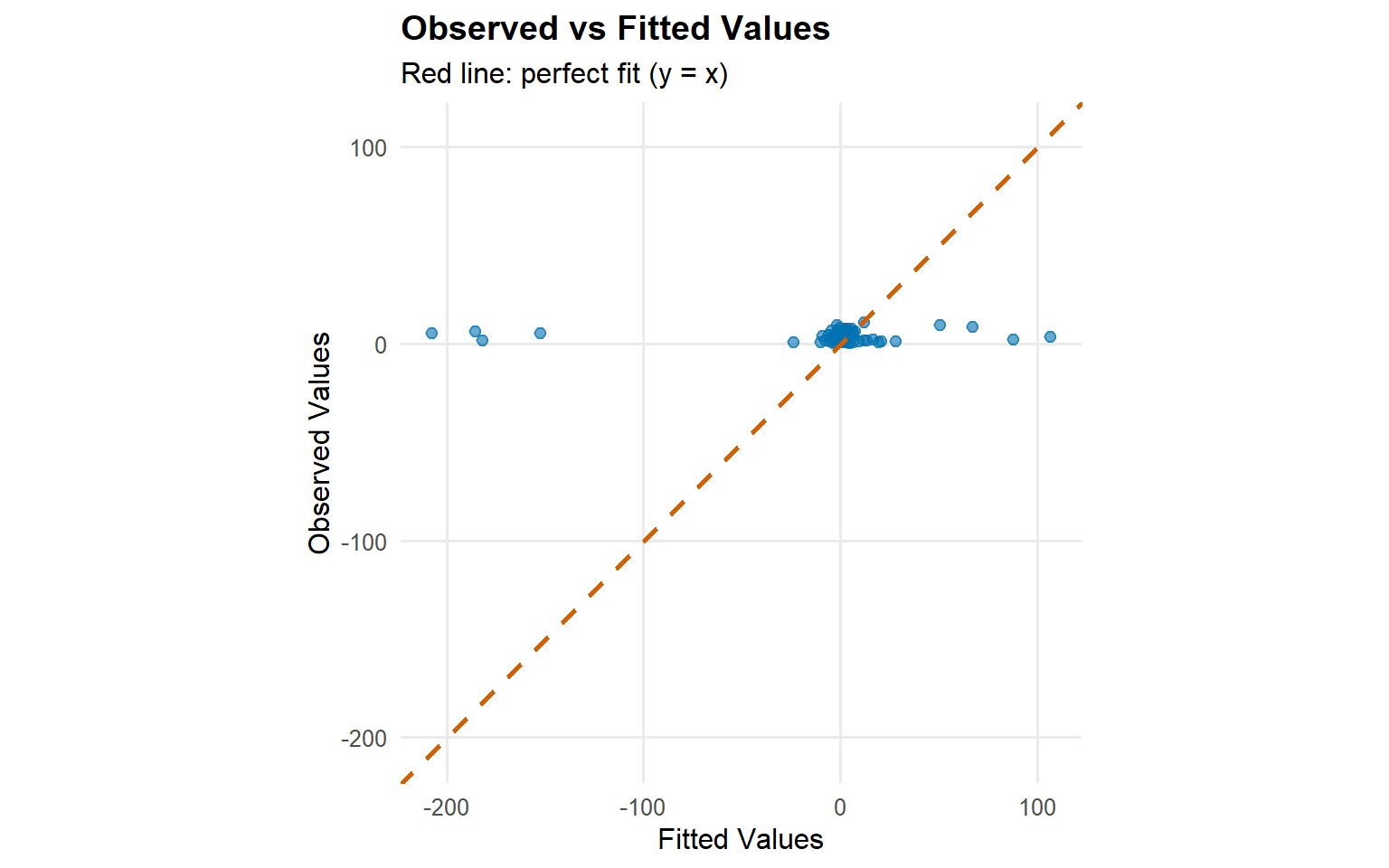
plot (fitted (fit_sb_cauchy))
Code
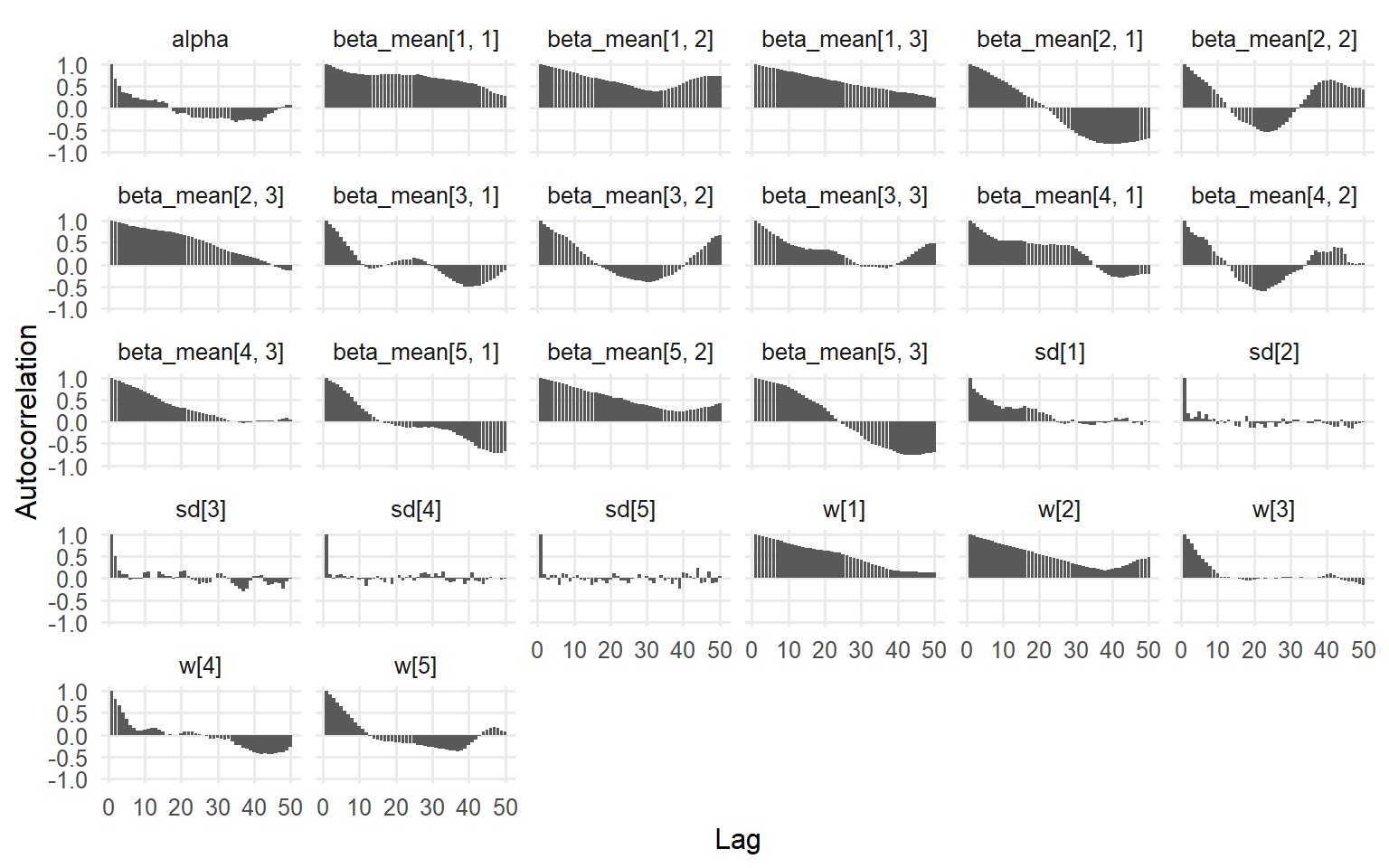
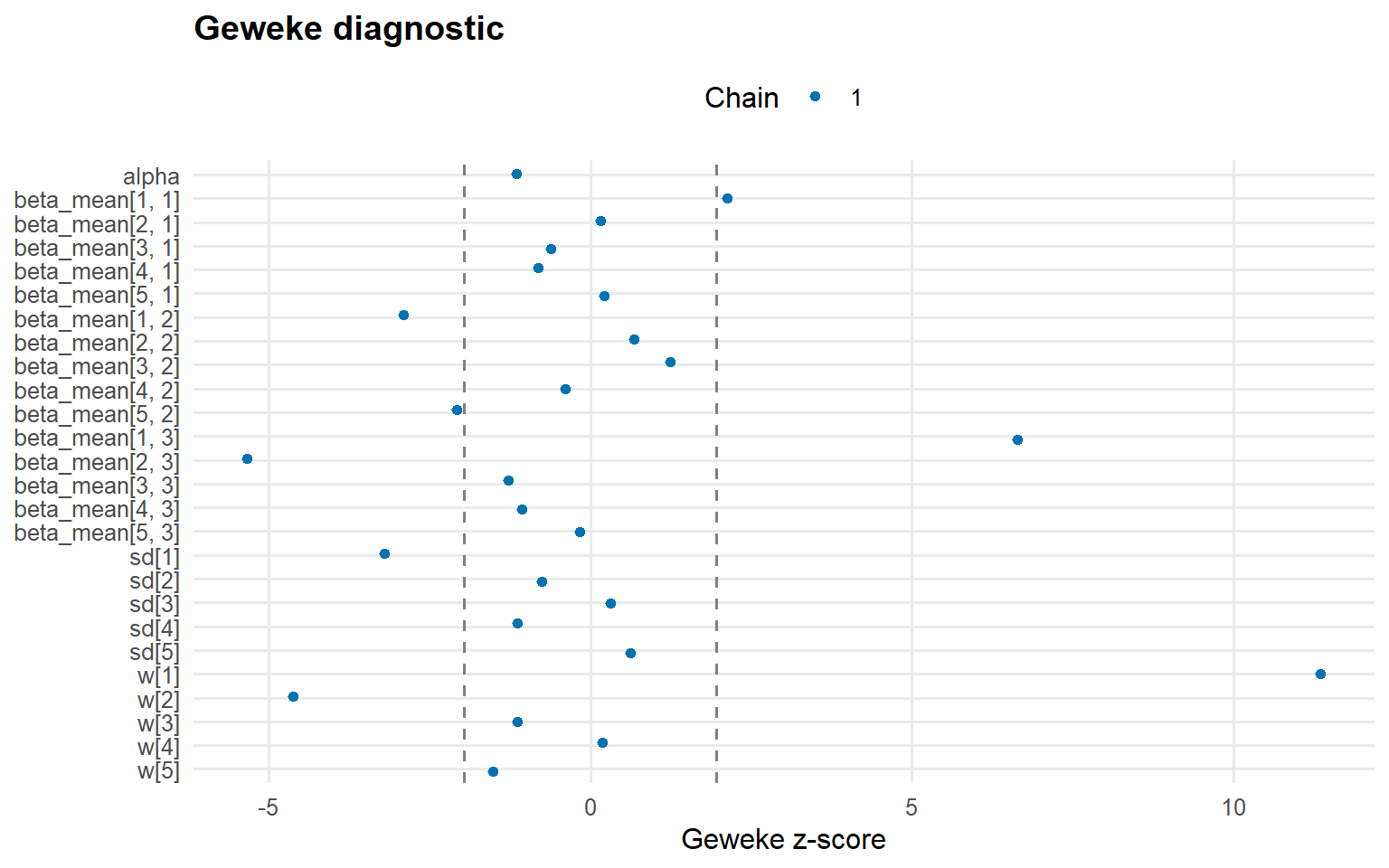
plot (fit_sb_normal, family = c ("traceplot" , "autocorrelation" , "geweke" ))
Code
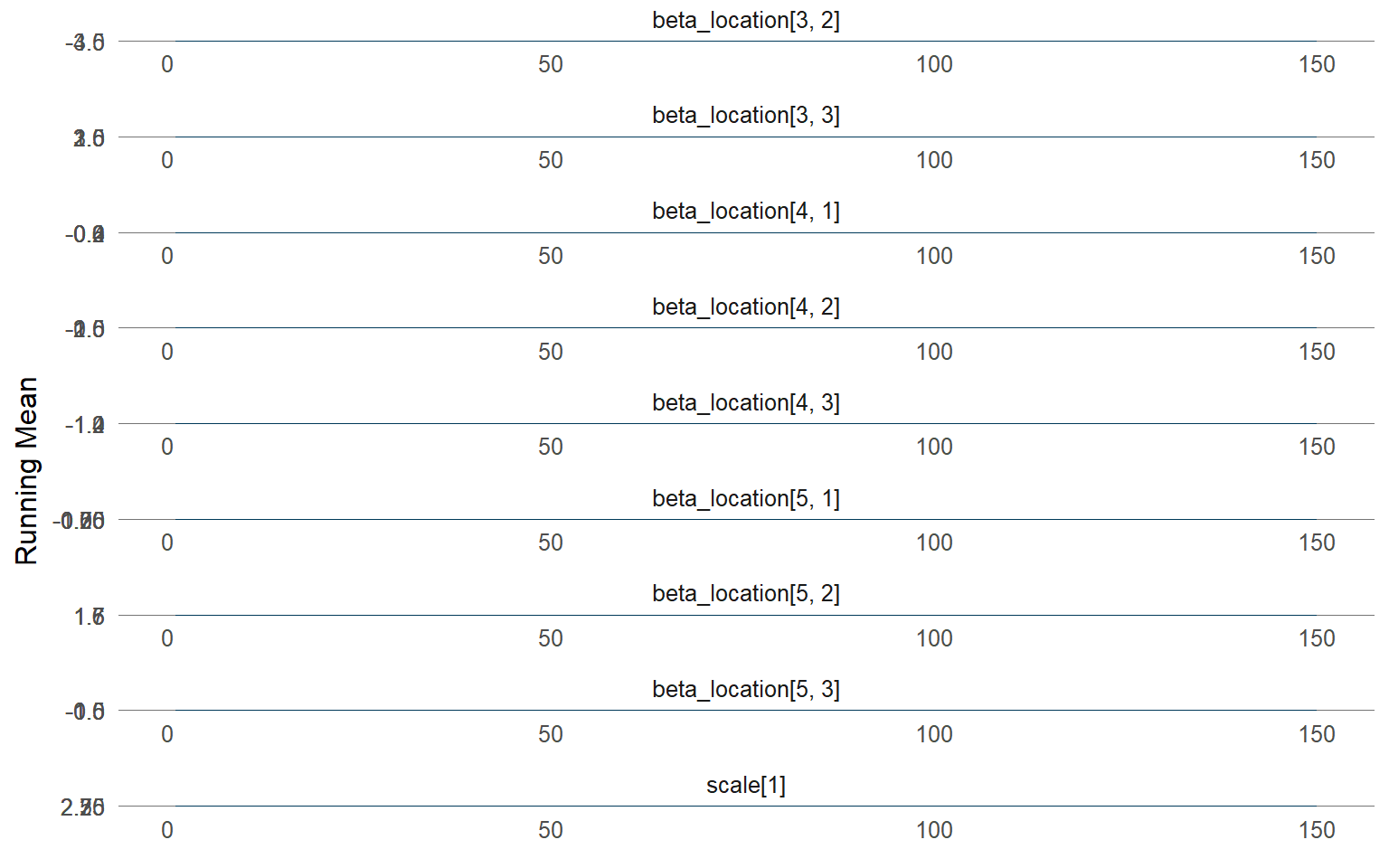
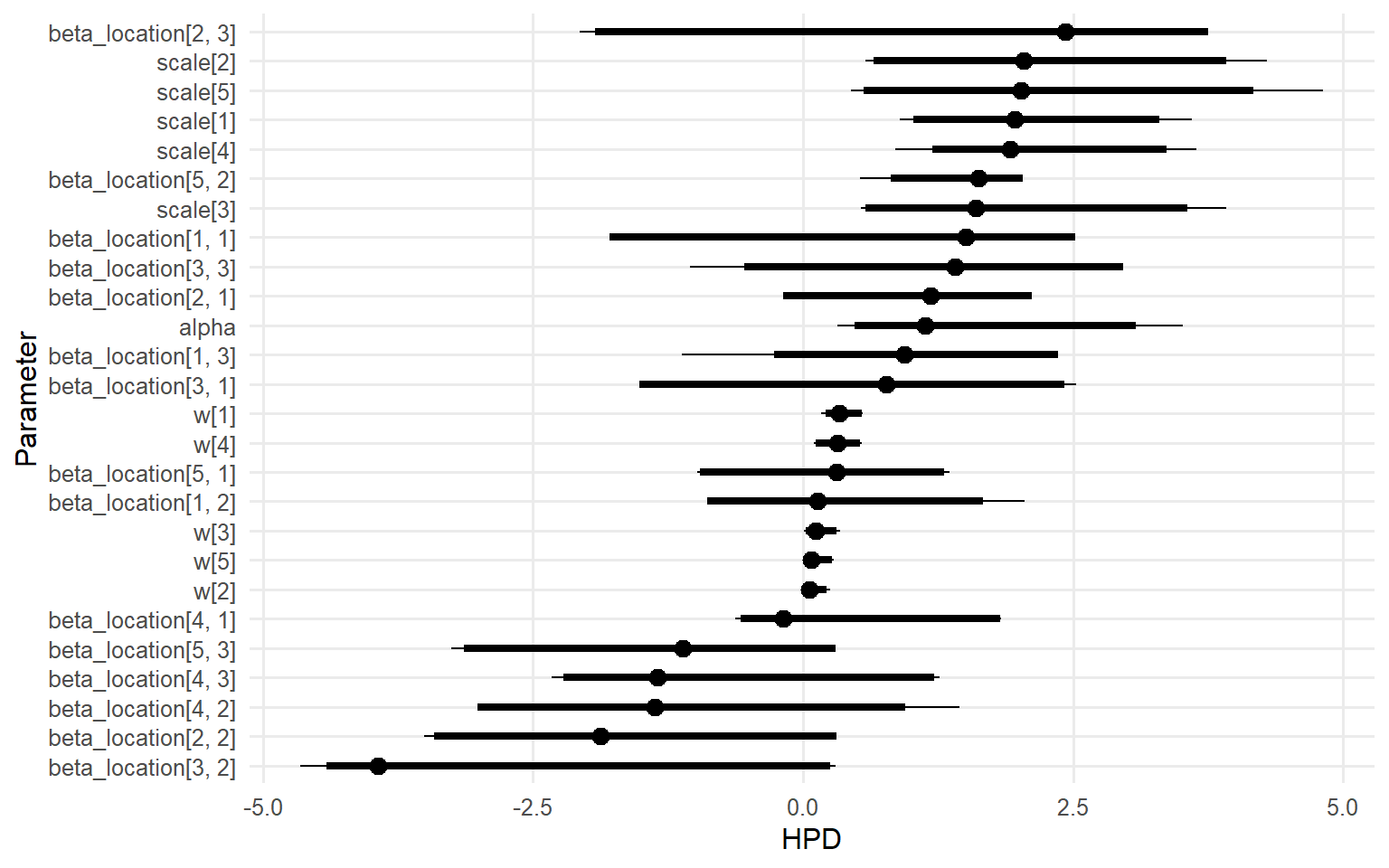
plot (fit_sb_cauchy, family = c ("density" , "running" , "caterpillar" ))
Takeaways
Stick-breaking component count is fixed but still supports covariate-dependent mixtures.
predict(..., type = "density") returns group-specific densities for each X.predict(..., type = "quantile") reports posterior-mean quantiles; the median is the 0.5 quantile and shifts with x1.Diagnoses rely on the same S3 plot()/fitted() pipeline available in other vignettes.
Prereqs
Required packages and data for this page are listed in the setup chunks above.
Outputs
This page renders model fits, diagnostics, and summary artifacts generated by package APIs.
Interpretation
Canonical concept page: Model Umbrella
Treat this page as an application/example view and use the canonical page for core definitions.
Next
Continue to the linked canonical concept page, then return for implementation-specific details.