X_new <- rbind(
c(-1, 0, 0, 0, 0),
c(0, 0, 0, 0, 0),
c(1, 1, 0, 0, 0)
)
colnames(X_new) <- colnames(X)
y_grid <- seq(0, max(y) * 1.2, length.out = 200)
df_pred_lognormal <- lapply(seq_len(nrow(X_new)), function(i) {
pred <- predict(fit_cond_gpd_lognormal, newdata =as.matrix(X_new[i, , drop = FALSE]), y = y_grid, type = "density")
data.frame(
y = pred$fit$y,
density = pred$fit$density,
label = paste("x1=", X_new[i, 1], ", x2=", X_new[i, 2], sep = ""),
model = "Lognormal"
)
})
df_pred_normal <- lapply(seq_len(nrow(X_new)), function(i) {
pred <- predict(fit_cond_gpd_normal, newdata =as.matrix(X_new[i, , drop = FALSE]), y = y_grid, type = "density")
data.frame(
y = pred$fit$y,
density = pred$fit$density,
label = paste("x1=", X_new[i, 1], ", x2=", X_new[i, 2], sep = ""),
model = "Normal"
)
})
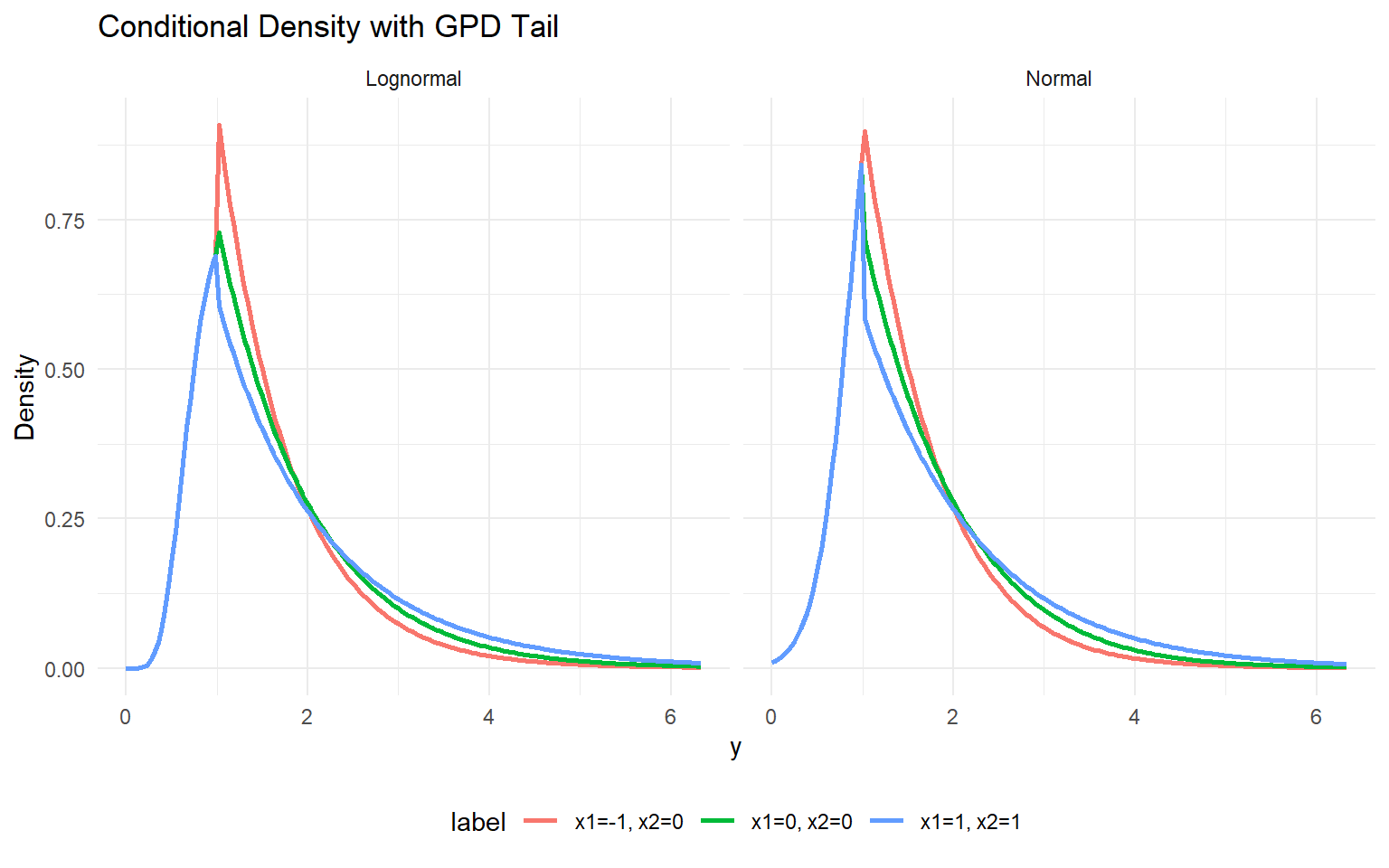
bind_rows(df_pred_lognormal, df_pred_normal) %>%
ggplot(aes(x = y, y = density, color = label)) +
geom_line(linewidth = 1) +
facet_wrap(~ model) +
labs(title = "Conditional Density with GPD Tail", x = "y", y = "Density") +
theme_minimal() +
theme(legend.position = "bottom")